primaryNeoplasmdp back to ToC or Data Property ToC
IRI: https://w3id.org/brainteaser/ontology/schema/primaryNeoplasm
- has domain
- ALS Brainteaser Comorbidities c
- has range
- boolean

This webpage describes the design and development of the BrainTeaser Ontology (BTO) whose purpose is to jointly model both Amyotrophic Lateral Sclerosis (ALS) and Multiple Sclerosis (MS) Clinical Data.
The BTO serves multiple purposes:
The BTO is innovative since it relies on very few seed concepts - Patient, Clinical Trial, Disease, Event - which allow us to jointly model ALS and MS and to grasp the time dimension entailed by the progression of such diseases.
Indeed, the core idea is that a Patient participates in a Clinical Trial, suffers from some Diseases, and undergoes Events. These Events are different in nature and cover a wide range of cases, e.g. Onset, Pregnancy, Symptom, Trauma, Diagnostic Procedure (like evoked potentials or ALS-FRS questionnaires) Therapeutic Procedure (like Mechanical Ventilation for ALS or Disease-Modifying Therapy for MS), Relapse, and more. Overall, this event-based approach allows us to model ALS and MS in an unified way, sharing concepts among these two diseases, and to track what happens during their progression. Details of the design and functioning of the BTO are provided in the next sections of this deliverable.
The BTO plays an important role in the overall BRAINTEASER architecture, shown in Figure 1. Indeed, it informs the implementation of the BRAINTEASER Semantic Data Cloud since the data contained here will be represented according to the BTO, i.e., they will be an instance of the BTO.
In Figure 1 it is possible to see that all the data will be anonymized prior to being represented in the BTO. This holds true for both the retrospective data, i.e., the data already held by clinical partners on the right of the figure, and the prospective data, i.e., the new data that will be collected during the project lifetime on the left.
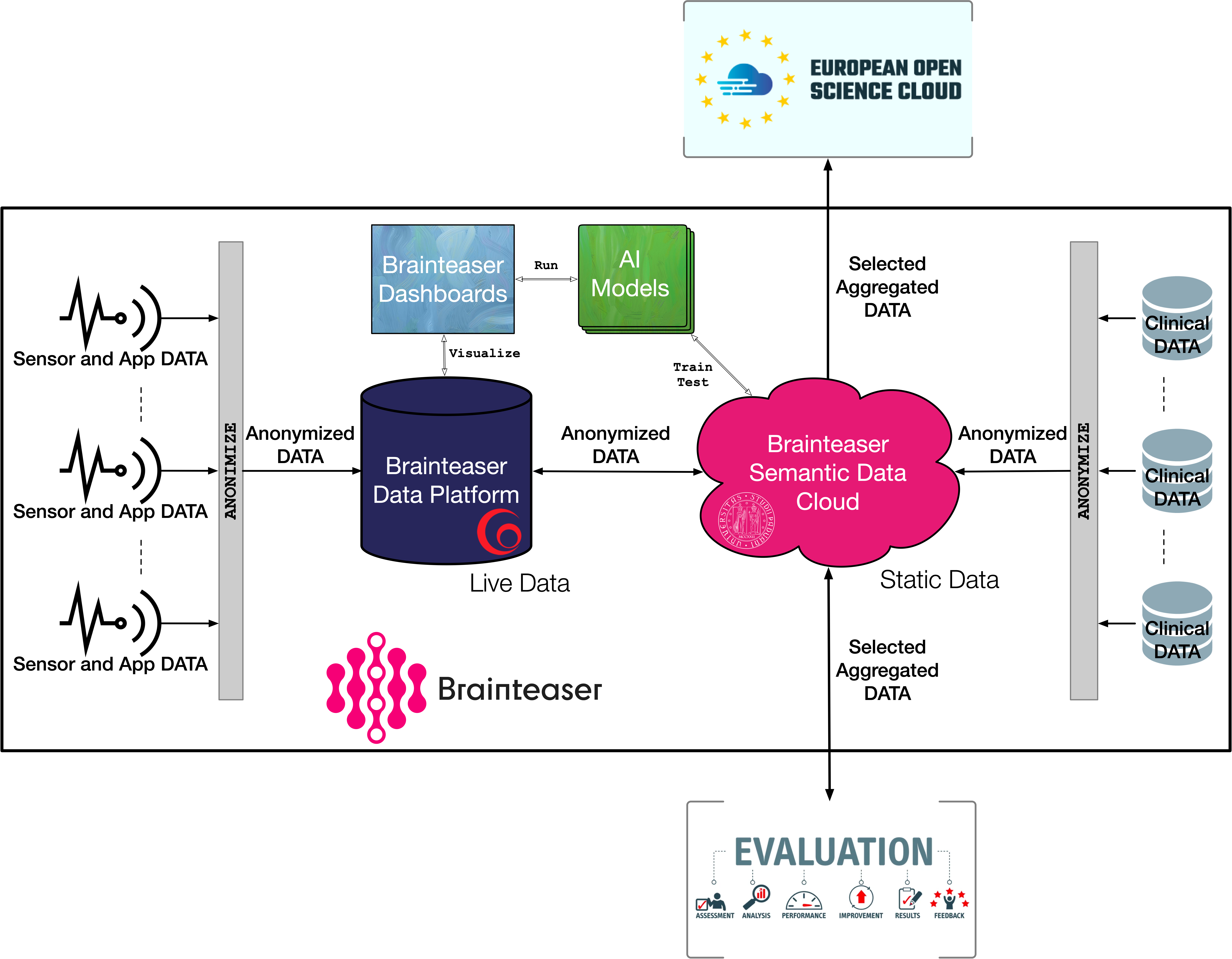
The data held in the BRAINTEASER Semantic Data Cloud, exported in a suitable format, will then be used to train the AI models needed to predict the progression of both ALS and MS.
Finally, a subset of the data in the BRAINTEASER Semantic Data Cloud will be exported to the European Open Science Cloud (EOSC) and they will be also shared and exploited for the Open Evaluation Challenges.
Figure 2 shows an overall picture of the whole data flow associated with the BTO and the BRAINTEASER Semantic Data Cloud, also in relation to Open Evaluation Challenges and the EOSC.
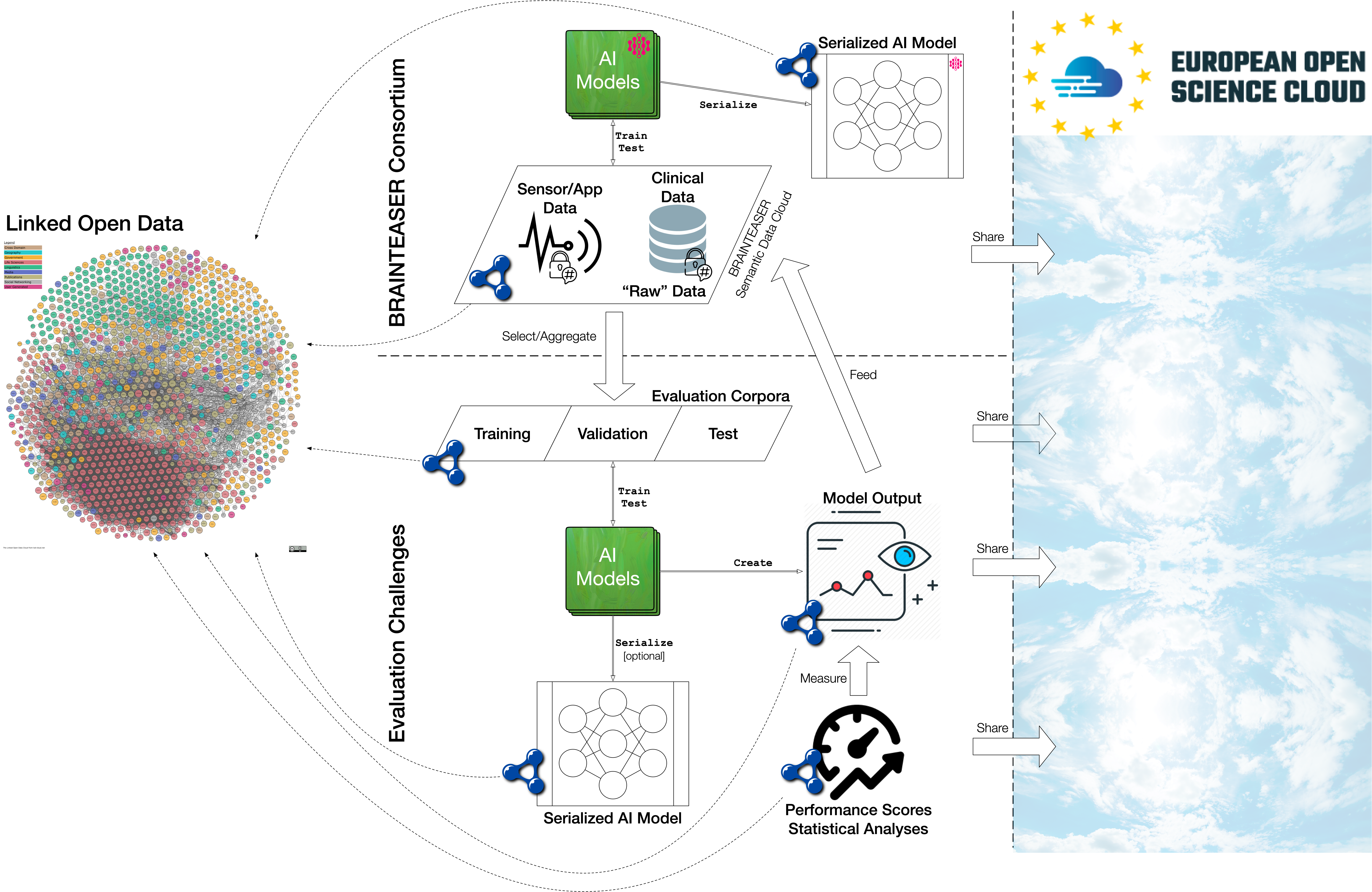
In summary, the BTO ontology will model the following data sources:
The BTO will also allow to link these data with other resources available in the Linked Open Data Cloud.
The term ontology has its origin in philosophy, where it refers to the study of existence. Similarly, in Computer Science an ontology is an explicit specification of a conceptualization [8]. Since their conception, ontologies have been largely employed to share a common understanding of the structure of information among people and software agents [9, 10]. On top of that, ontologies allow computers to have access to structured collections of information and rules that can be used to conduct automated reasoning. For these reasons, ontologies are considered the third basic component of the Semantic Web [9, 11]. Depending on the generality of the instances included within ontologies, we can distinguish between foundation and domain ontologies. The former model com- mon concepts that can be used in a wide variety of domains. A notable example is the Friend-Of-A-Friend (FOAF) ontology, which describes people’s personal information as well as their social relationships [11]. In general, foundation ontologies are thought to be imported and reused in domain-specific applications, where the original components are refined and extended to better represent the modeled reality [10]. The latter model a specific domain, thus representing the particular meaning of terms as they apply to that specialization. Indeed, the same word can assume different meanings in different fields. To illustrate this, let’s take the word “polyp” as an example. In the medical domain, a polyp is an abnormal growth of tissue projecting from a mucous membrane. On the other hand, in zoology, a polyp represents one of the two forms found in the Phylum Cnidaria. Given the objectives of this work, below we focus on describing domain ontologies.
External Referencing Whenever we model clinical information, it is preferable to use external ontologies to provide correspondence between largely used medical terminology. Due to its widely diffused adoption and exhaustiveness, our primary taxonomy of choice is NCIT [30], but we also employed other well-known resources when no information is available in NCIT, e.g. SNOMED-CT or ATC. In general, this is common practice when developing ontologies [44]. In fact, by reusing entities and properties already defined in other ontologies, we are not only able to enforce collaboration and data consistency among databases but to guarantee the authori- tativeness of the semantic meaning of these resources. External referencing has been managed with annotation properties and using the URI of the term in the original taxonomy. In general, we used external URIs when we defined named individuals that refer to abstract concepts. On the contrary, when we inserted a new class in BTO we used the Brainteaser namespace and connected references using annotation properties.
Namespaces BTO URIs are divided into two namespaces: the schema namespace https://w3id.org/brainteaser/ontology/schema/ and the resource namespace https://w3id.org/brainteaser/ontology/resource/. All URIs corresponding to classes, data properties, and object properties belong to the former namespace. At the same time, in the latter, we include all URIs referring to real-world instances of the entities described in BTO at an ontological level. Notice that, in this sense, the resource namespace is empty until the practitioner starts populating it with real-world data. The only instances included in the schema namespace are the named individuals corresponding to Simple Knowledge Organization System (SKOS) con- cepts (as defined in Subsection 3.4). Our choice of including these elements in the schema namespace stems from the fact that, akin to relational modeling controlled dictionaries, these entities do not depend on the data underneath but can be seen as a predefined thesaurus of concepts and are a fundamental part of the environment that we model.
Classes Definition and Annotation Properties All components of the BTO have additional information in the form of annotation properties. We defined a list of essential metadata to add when we introduced a new class. Firstly, all classes must have a label denoting the name and a comment, which provides a brief explana- tion together with its source. If the class had an equivalent in NCIT, we inherited the name and definition. In this case, we also added another annotation property called “rdfs:isDefinedBy with the Internationalized Resource Identifier (IRI) corre- sponding to the NCIT term of reference. Most biomedical vocabularies are mapped in the UMLS with a unique identifier called Concept Unique Identifier (CUI) [40]. For each class that had a UMLS reference, we added the annotation property “dcterms:conformsTo” with the URL of the corresponding concept.
Naming Conventions Regarding object and data properties, we applied a similar approach and defined the needed annotations and naming conventions to provide uniformity throughout BTO. In particular, all components must have a label and a comment. About object properties, we used explanatory labels where we included the property range. In this case, the comment explains the relationship between the two classes involved. Concerning data properties, the label usually includes the name of the domain class so that its meaning is intuitive. We also added a comment with the attribute description and, when available, the definition source. In conclusion, we defined all data properties to achieve consistency when specifying datatypes. For example, all string properties have datatype “rdf:PlainLiteral”. We also used the “note” annotation property in all BTO components for additional remarks or business logic rules.
| Class | Reference Taxonomy |
|---|---|
| Kinship Type | National Cancer Institute Thesaurus (NCIT) |
| Occupation | Occupations pillar of the ESCO Classification (ESCO) |
| Group (social concept) | SNOMED Clinical Terms (SNOMED-CT) |
| Gene | National Cancer Institute Thesaurus (NCIT) |
| Disease, Disorder or Findings | National Cancer Institute Thesaurus (NCIT) |
| Surgical Procedure Type | SNOMED Clinical Terms (SNOMED-CT) |
| Therapeutic Procedure Type | National Cancer Institute Thesaurus (NCIT) |
| Anatomic Site | National Cancer Institute Thesaurus (NCIT) |
| Pharmacological Substance | Anatomical Therapeutic Chemical (ATC) Classification |
| Advesre Drug Reaction | Ontology of Adverse Events (OAE) |

Figure 3: Patient semantic area. “Patient” is a subclass of the class Person from FOAF ontology. We directly connect to patient data about genetic mutations, occupation, family history, and residence place. The latter is useful to link environmental information to each patient, which is modeled by importing the “Global City Indicator Environment Ontology” [38].
Environmental Data
Besides genetic predisposition, lifestyle and environmental factors are shown to
affect the risk of MS. Once we acknowledge such correlation, some of these environmental
and lifestyle factors can be modified with prevention measures, especially
for people at greater risk [45]. On the other hand, identifying environmental factors
linked to ALS has proven to be challenging and it still lacks large cohorts. In both
cases, understanding the interaction between genetic and environmental factors can
be a great resource for integrating precision medicine into ALS or MS care [46].
That is why connecting environmental data to each patient in BTO is a fundamental
requirement. These environmental data include pollution information for a given
location, recorded by a weather station. To store this piece of information, in BTO
the practitioner can record each patient’s birth and residence places using the class
“Place”. To model environmental data, we imported Global City Indicator Environment
Ontology[38], also known as Pollution ontology, an ontology designed to
represent ontologically environmental data registered by weather stations. In par-
ticular, we connected a weather station, modeled with the class “Sensing Device”
from the above-mentioned ontology, to a location, modeled with the class “Place”,
with the object property “:coveredPlace”, to model the fact that a station registers
environmental data for a given location. We report how we integrated air pollutants in
Figure 4. The concentration of each air pollutant of interest is
modeled as a subclass of “Air Pollution Concentration”. Notice that we are interested in
storing information about Particulate Matter (PM) 10, PM 2.5, Ozone (O3), Nitrogen
Dioxide (NO2), Sulphur Dioxide (SO2), and Carbon Monoxide (CO) thus we
created a class in our namespace for pollutants not represented in the imported
ontology, such as CO. Air pollution concentrations are produced by sensing de-
vices, thus we connected the classes “Air Pollution Concentration” and “Sensing
Device” with the property “ssn:isProducedBy”. The concentration value of
each air pollutant is stored using the data property “numerical value”, while
the date of the measure is referred to with the object property ":APConcentrationTIme" that links
“Air Pollution Concentration” class to class “time:Instant”. All classes, object
properties, and data properties mentioned above are imported from the “Global
City Indicator Environment Ontology”[38].
Figure 4: Environmental data integration using the imported “Global City Indicator
Environment Ontology” [38]. We linked the class “Place” with “Sensing
Device”, which produces pollution concentration data. Such information
is stored using one of the subclasses of “Air Pollution Concentration”, which
represents the specific pollutant.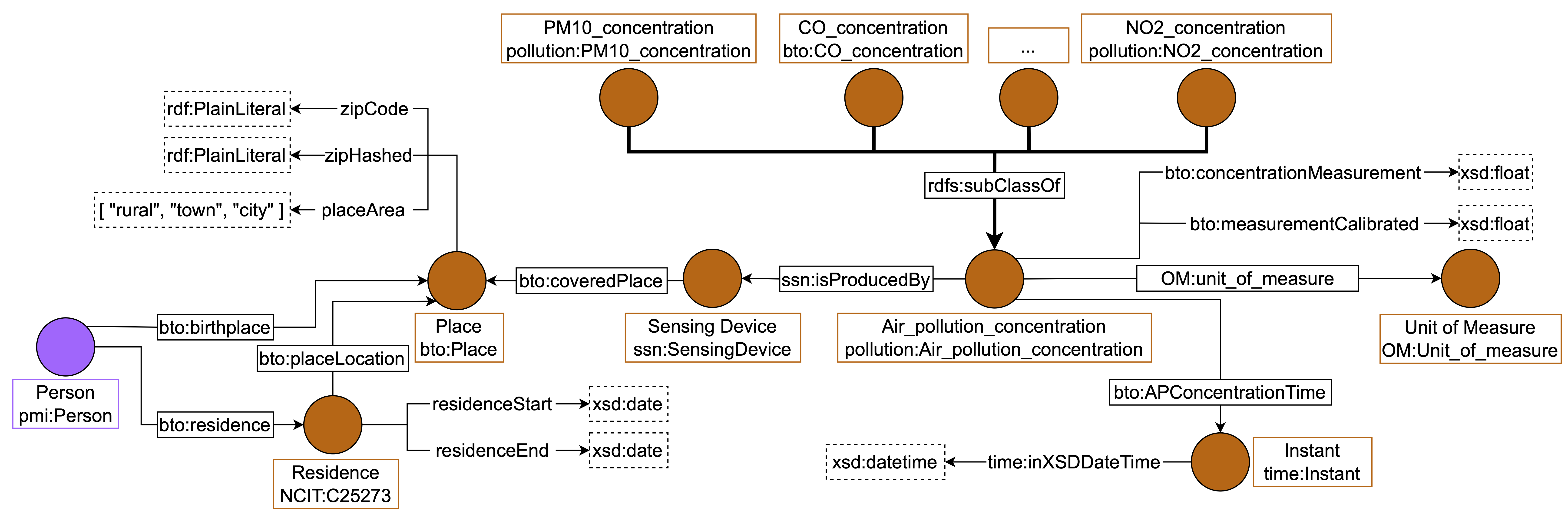

Figure 5: Event semantic area. Each patient is connected to clinical history and lifestyle information through the class Event. In fact, using such nodes physicians can store data about habits, past traumas, any clinical event (class “Condition”), coexisting medical conditions (called “comorbidities”), or pregnancies and MS patients’ relapses.
Disease, Disorder or Finding Taxonomy
The class “Disease, Disorder or Finding” includes diseases, like carcinoma or chickenpox infection, but also injuries, symptoms, and findings. This allows modeling and storing in a KB any sort of clinical event that occurred to a patient, even if it is not directly linked to ALS or MS. We modeled this class following the SKOS data model, using NCIT as reference taxonomy, with a few additions from SNOMED-CT thesaurus. As illustrated in Figure 5, we modeled past traumas with the class “Trauma”. However, some injuries have also been made available in the “Disease, Disorder or Finding” taxonomy. As a general rule, we use the taxonomy whenever the trauma is specific (e.g., “Open Dislocation of the Shoulder”). On the other hand, if the clinician knows only that a patient suffered from generic trauma to the shoulder, we instantiate an individual of type “Trauma” and connect it to its Anatomic Site (e.g., “shoulder”) with the property “:anatomicalLocation”. Notice that, thanks to SKOS characteristics, specific traumas (e.g., “open dislocation of the shoulder”) are in relation “:hasBroaderTransitive” with the general concept “injury”. We follow a similar line of thought to deal with symptoms. In this case, we use the taxonomy to model cases where the practitioner needs to store information about general symptoms (e.g., headache, fever). A particular group of symptoms that required deeper modeling granularity is the one that constitutes the onset of the disease (either ALS or MS). Given the high relevance that such symptoms have on the course of the disease, they have not been modeled using the symptoms taxonomy but as data properties of the class “Onset”.
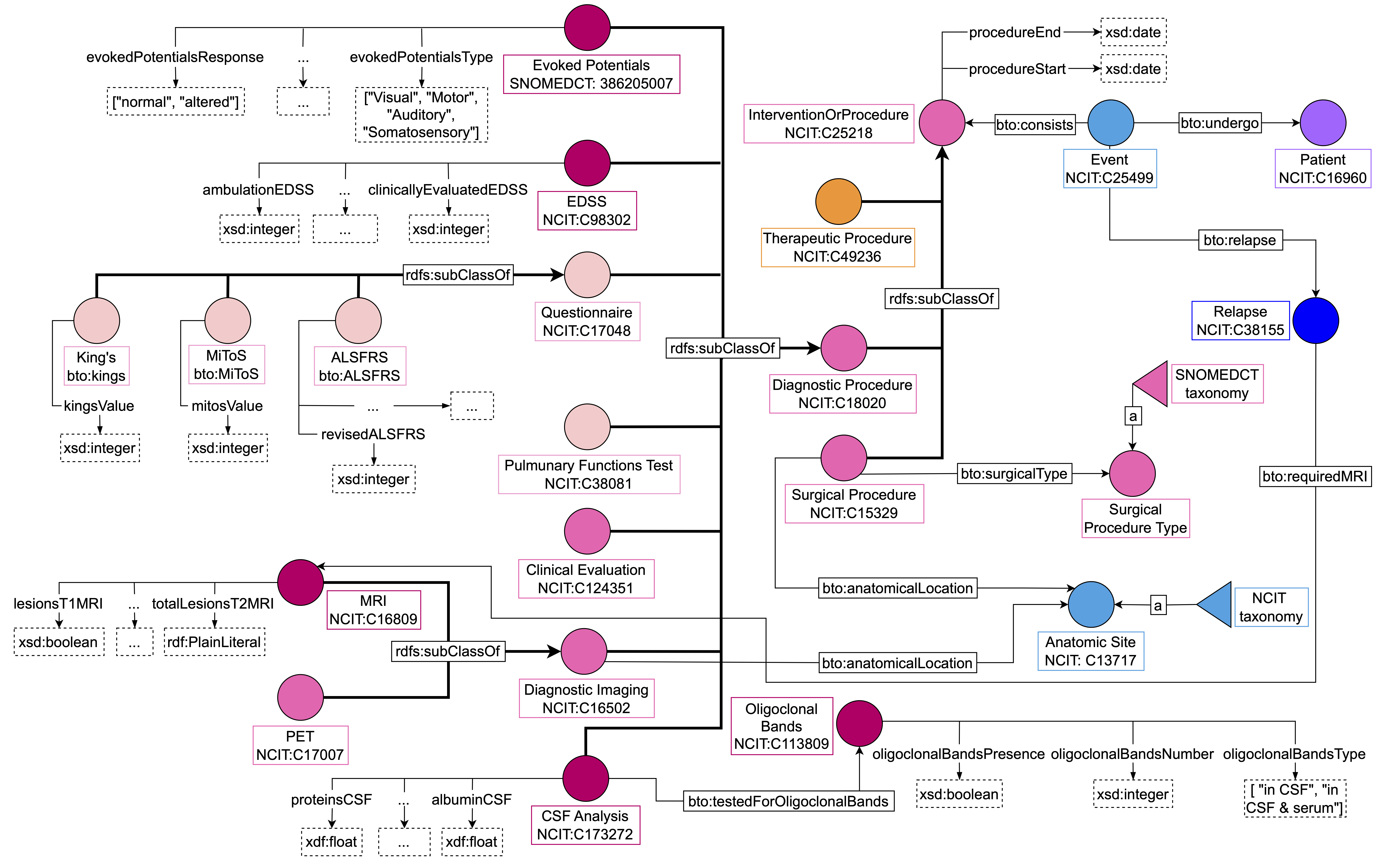
Figure 6: “Intervention or Procedures” area. We distinguish between surgical procedures, diagnostic procedures, and therapeutic procedures. Both surgical and therapeutic interventions are the same for MS and ALS patients while diagnostic procedures can differ based on the disease.

Figure 7: “Therapeutic Procedures” area. The core of the area is the “Therapeutic Treatment” class, where one can store information about drug administrations and its specific aspects, such as dose, frequency and end treatment. In particular, we modeled the “End Therapy Reason” as a class so that we can link possible causes, such as adverse events and pregnancy.
IRI: https://w3id.org/brainteaser/ontology/schema/Activity
IRI: https://w3id.org/brainteaser/ontology/schema/Administration
IRI: https://w3id.org/brainteaser/ontology/schema/AdverseDrugReaction
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#Air_pollution
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#Air_pollution_concentration
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRS
IRI: https://w3id.org/brainteaser/ontology/schema/AnatomicalSite
IRI: https://w3id.org/brainteaser/ontology/schema/AtmosphericPressure
IRI: https://w3id.org/brainteaser/ontology/schema/AwajiClinicalEvaluation
IRI: https://w3id.org/brainteaser/ontology/schema/BeatToBeatData
IRI: https://w3id.org/brainteaser/ontology/schema/BeforeOnset
IRI: https://w3id.org/brainteaser/ontology/schema/C6H6_concentration
IRI: https://w3id.org/brainteaser/ontology/schema/BVMTR
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#CH4
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#CH4_concentration
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#CO2
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#CO2_concentration
IRI: https://w3id.org/brainteaser/ontology/schema/CVLTII
IRI: https://w3id.org/brainteaser/ontology/schema/CO_concentration
IRI: https://w3id.org/brainteaser/ontology/schema/ClinicalEvaluation
IRI: https://w3id.org/brainteaser/ontology/schema/ClinicalTrial
IRI: https://w3id.org/brainteaser/ontology/schema/ClinicalTrialParticipation
IRI: https://w3id.org/brainteaser/ontology/schema/ClinicsAndHospitals
IRI: https://w3id.org/brainteaser/ontology/schema/CognitiveAssessment
IRI: http://www.w3.org/2004/02/skos/core#Collection
IRI: https://w3id.org/brainteaser/ontology/schema/Comorbidity
IRI: http://www.wurvoc.org/vocabularies/om-1.8/Compount_unit
IRI: http://www.w3.org/2004/02/skos/core#Concept
IRI: http://www.w3.org/2004/02/skos/core#ConceptScheme
Thesauri, classification schemes, subject heading lists, taxonomies, 'folksonomies', and other types of controlled vocabulary are all examples of concept schemes. Concept schemes are also embedded in glossaries and terminologies.
IRI: https://w3id.org/brainteaser/ontology/schema/Condition
IRI: https://w3id.org/brainteaser/ontology/schema/CSFAnalysis
IRI: http://www.wurvoc.org/vocabularies/om-1.8/cubic_metre
IRI: https://w3id.org/brainteaser/ontology/schema/AvgRelHumidity
IRI: https://w3id.org/brainteaser/ontology/schema/AvgSeaLvlPressure
IRI: https://w3id.org/brainteaser/ontology/schema/MaxTemp
IRI: https://w3id.org/brainteaser/ontology/schema/MeanGlobRadiation
IRI: https://w3id.org/brainteaser/ontology/schema/MeanTemp
IRI: https://w3id.org/brainteaser/ontology/schema/MeanWindSpeed
IRI: https://w3id.org/brainteaser/ontology/schema/MinTemp
IRI: https://w3id.org/brainteaser/ontology/schema/PrecipitationSum
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#decibel
IRI: https://w3id.org/brainteaser/ontology/schema/Diagnosis
IRI: https://w3id.org/brainteaser/ontology/schema/DiagnosticImaging
IRI: https://w3id.org/brainteaser/ontology/schema/DiagnosticProcedure
IRI: https://w3id.org/brainteaser/ontology/schema/DiseaseDisorderOrFinding
IRI: https://w3id.org/brainteaser/ontology/schema/DYMUS
IRI: https://w3id.org/brainteaser/ontology/schema/ECAS
IRI: https://w3id.org/brainteaser/ontology/schema/EndTherapyReason
IRI: https://w3id.org/brainteaser/ontology/schema/EQ5D5L
IRI: https://w3id.org/brainteaser/ontology/schema/Event
IRI: https://w3id.org/brainteaser/ontology/schema/EvokedPotentials
IRI: https://w3id.org/brainteaser/ontology/schema/EDSS
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#GHGs
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#GHGs_concentration
IRI: https://w3id.org/brainteaser/ontology/schema/Gene
IRI: https://w3id.org/brainteaser/ontology/schema/GeneticTesting
IRI: https://w3id.org/brainteaser/ontology/schema/GlobalRadiation
IRI: https://w3id.org/brainteaser/ontology/schema/GoldCoastCriteria
IRI: https://w3id.org/brainteaser/ontology/schema/Group
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#HFCs
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#HFCs_concentration
IRI: https://w3id.org/brainteaser/ontology/schema/HandDexterity
IRI: https://w3id.org/brainteaser/ontology/schema/HeartRateMeasurement
IRI: https://w3id.org/brainteaser/ontology/schema/HematologyTest
IRI: https://w3id.org/brainteaser/ontology/schema/HADS
IRI: https://w3id.org/brainteaser/ontology/schema/HADS_ALS
IRI: https://w3id.org/brainteaser/ontology/schema/Humidity
IRI: https://w3id.org/brainteaser/ontology/schema/Impairment
IRI: http://www.w3.org/2006/time#Instant
IRI: http://www.w3.org/2006/time#Interval
IRI: https://w3id.org/brainteaser/ontology/schema/InterventionOrProcedure
IRI: https://w3id.org/brainteaser/ontology/schema/KingsClinicalStaging
IRI: https://w3id.org/brainteaser/ontology/schema/Kinship
IRI: https://w3id.org/brainteaser/ontology/schema/KinshipType
IRI: https://w3id.org/brainteaser/ontology/schema/LaboratoryTestResult
IRI: https://w3id.org/brainteaser/ontology/schema/Last5YearsEvent
IRI: https://w3id.org/brainteaser/ontology/schema/Lifestyle
IRI: https://w3id.org/brainteaser/ontology/schema/MRI
IRI: http://www.wurvoc.org/vocabularies/om-1.8/Mass
IRI: https://w3id.org/brainteaser/ontology/schema/MenstrualCycle
IRI: http://www.wurvoc.org/vocabularies/om-1.8/microgram
IRI: http://ontology.eil.utoronto.ca/GCI/Foundation/GCI-Foundation.owl#microgram_per_cubic_metre
IRI: https://w3id.org/brainteaser/ontology/schema/MiToSFunctionalStaging
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS
IRI: https://w3id.org/brainteaser/ontology/schema/MoreThan5YearsEvent
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54
IRI: https://w3id.org/brainteaser/ontology/schema/MuscleToneALS
IRI: https://w3id.org/brainteaser/ontology/schema/MuscleToneAssessment
IRI: https://w3id.org/brainteaser/ontology/schema/MuscleToneMS
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#NO2
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#N2O
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#N2O_concentration
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#NO2_concentration
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#Noise_pollution
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#Noise_pollution_concentration
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#O3
IRI: https://w3id.org/brainteaser/ontology/schema/Occupation
IRI: https://w3id.org/brainteaser/ontology/schema/OligoclonalBands
IRI: https://w3id.org/brainteaser/ontology/schema/Onset
IRI: http://www.w3.org/2004/02/skos/core#OrderedCollection
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#O3_concentration
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#PFCs
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#PFCs_concentration
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#PM
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#PM10
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#PM10_Mass
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#PM10_Volume
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#PM2.5
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#PM2.5_Mass
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#PM2.5_Volume
IRI: https://w3id.org/brainteaser/ontology/schema/PM1_concentration
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#PM10_concentration
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#PM2.5_concentration
IRI: https://w3id.org/brainteaser/ontology/schema/Patient
IRI: http://www.w3.org/2000/10/swap/pim/contact#Person
IRI: https://w3id.org/brainteaser/ontology/schema/PharmacologicSubstance
IRI: https://w3id.org/brainteaser/ontology/schema/PhysicalActivity
IRI: https://w3id.org/brainteaser/ontology/schema/PhysicalActivityData
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI
IRI: https://w3id.org/brainteaser/ontology/schema/Place
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#Pollution
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#Pollution_average
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#Pollution_concentration
IRI: https://w3id.org/brainteaser/ontology/schema/PET
IRI: https://w3id.org/brainteaser/ontology/schema/Precipitation
IRI: https://w3id.org/brainteaser/ontology/schema/Pregnancy
IRI: https://w3id.org/brainteaser/ontology/schema/ProtocolEvent
IRI: https://w3id.org/brainteaser/ontology/schema/PulmonaryFunctionTest
IRI: https://w3id.org/brainteaser/ontology/schema/PFTs
IRI: http://www.wurvoc.org/vocabularies/om-1.8/Quantity
IRI: https://w3id.org/brainteaser/ontology/schema/Questionnaire
IRI: https://w3id.org/brainteaser/ontology/schema/AppQuestionnaire
IRI: https://w3id.org/brainteaser/ontology/schema/Relapse
IRI: https://w3id.org/brainteaser/ontology/schema/Residence
IRI: https://w3id.org/brainteaser/ontology/schema/RespiratoryData
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#SF6
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#SF6_concentration
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#SO2
IRI: https://w3id.org/brainteaser/ontology/schema/SelfALSFRSR
IRI: http://purl.oclc.org/NET/ssnx/ssn#SensingDevice
IRI: http://purl.oclc.org/NET/ssnx/ssn#Sensor
IRI: http://www.wurvoc.org/vocabularies/om-1.8/Singular_unit
IRI: https://w3id.org/brainteaser/ontology/schema/SleepData
IRI: https://w3id.org/brainteaser/ontology/schema/SLEEP_ALS
IRI: https://w3id.org/brainteaser/ontology/schema/Smoking
IRI: https://w3id.org/brainteaser/ontology/schema/SPO2Data
IRI: https://w3id.org/brainteaser/ontology/schema/StressLevel
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#SO2_concentration
IRI: https://w3id.org/brainteaser/ontology/schema/SurgicalProcedure
IRI: https://w3id.org/brainteaser/ontology/schema/SurgicalType
IRI: https://w3id.org/brainteaser/ontology/schema/SDMT
IRI: https://w3id.org/brainteaser/ontology/schema/Symptom
IRI: https://w3id.org/brainteaser/ontology/schema/Temperature
IRI: https://w3id.org/brainteaser/ontology/schema/TherapeuticProcedure
IRI: https://w3id.org/brainteaser/ontology/schema/TherapeuticProcedureType
IRI: https://w3id.org/brainteaser/ontology/schema/TherapeuticTreatment
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#tonne
IRI: https://w3id.org/brainteaser/ontology/schema/Trauma
IRI: http://www.wurvoc.org/vocabularies/om-1.8/Unit_division
IRI: http://www.wurvoc.org/vocabularies/om-1.8/Unit_multiple_or_submultiple
IRI: http://www.wurvoc.org/vocabularies/om-1.8/Unit_of_measure
IRI: https://w3id.org/brainteaser/ontology/schema/UseCaseComorbidities
IRI: https://w3id.org/brainteaser/ontology/schema/VisualAcuity
IRI: https://w3id.org/brainteaser/ontology/schema/VAS
IRI: https://w3id.org/brainteaser/ontology/schema/VAS_ALS
IRI: https://w3id.org/brainteaser/ontology/schema/VOC_concentration
IRI: http://www.wurvoc.org/vocabularies/om-1.8/Volume
IRI: https://w3id.org/brainteaser/ontology/schema/WalkTest
IRI: https://w3id.org/brainteaser/ontology/schema/WalkTestALS
IRI: https://w3id.org/brainteaser/ontology/schema/WalkTestMS
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#Water_pollution
IRI: https://w3id.org/brainteaser/ontology/schema/WearableDataMeasurement
IRI: https://w3id.org/brainteaser/ontology/schema/WeatherData
IRI: https://w3id.org/brainteaser/ontology/schema/WEIGHT
IRI: https://w3id.org/brainteaser/ontology/schema/WindSpeed
IRI: https://w3id.org/brainteaser/ontology/schema/administration
IRI: https://w3id.org/brainteaser/ontology/schema/adverseReaction
IRI: https://w3id.org/brainteaser/ontology/schema/anatomicalLocation
IRI: https://w3id.org/brainteaser/ontology/schema/APConcentrationTime
IRI: https://w3id.org/brainteaser/ontology/schema/birthplace
IRI: https://w3id.org/brainteaser/ontology/schema/coexistingPregnancy
IRI: https://w3id.org/brainteaser/ontology/schema/conductedInClinic
IRI: https://w3id.org/brainteaser/ontology/schema/confirmedPregnancy
IRI: https://w3id.org/brainteaser/ontology/schema/consists
IRI: https://w3id.org/brainteaser/ontology/schema/coveredPlace
IRI: https://w3id.org/brainteaser/ontology/schema/determinedBy
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/enrolledIn
IRI: https://w3id.org/brainteaser/ontology/schema/filledAppQuestionnaire
IRI: http://www.w3.org/2004/02/skos/core#broader
IRI: http://www.w3.org/2004/02/skos/core#broadMatch
IRI: http://www.w3.org/2004/02/skos/core#broaderTransitive
has characteristics: transitive
IRI: http://www.w3.org/2004/02/skos/core#closeMatch
has characteristics: symmetric
IRI: http://www.w3.org/2004/02/skos/core#exactMatch
has characteristics: symmetric, transitive
IRI: http://www.w3.org/2004/02/skos/core#member
IRI: http://www.w3.org/2004/02/skos/core#memberList
has characteristics: functional
IRI: http://www.w3.org/2004/02/skos/core#narrower
IRI: http://www.w3.org/2004/02/skos/core#narrowMatch
IRI: http://www.w3.org/2004/02/skos/core#narrowerTransitive
has characteristics: transitive
IRI: http://www.w3.org/2004/02/skos/core#hasTopConcept
IRI: https://w3id.org/brainteaser/ontology/schema/hasDisease
IRI: https://w3id.org/brainteaser/ontology/schema/hasEthnicity
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/hasKinship
IRI: https://w3id.org/brainteaser/ontology/schema/hasKinshipType
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/hasOccupation
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/hasRegisteredActivity
IRI: https://w3id.org/brainteaser/ontology/schema/hasRegisteredCondition
IRI: http://www.w3.org/2006/time#inDateTime
IRI: https://w3id.org/brainteaser/ontology/schema/inClinicalTrial
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/inKinshipWith
IRI: http://www.w3.org/2004/02/skos/core#mappingRelation
IRI: http://www.w3.org/2004/02/skos/core#inScheme
IRI: http://www.w3.org/2004/02/skos/core#semanticRelation
IRI: http://purl.oclc.org/NET/ssnx/ssn#isProducedBy
IRI: http://www.w3.org/2004/02/skos/core#topConceptOf
IRI: https://w3id.org/brainteaser/ontology/schema/isAboutDisease
IRI: http://ontology.eil.utoronto.ca/GCI/Environment/Pollution.owl#measures
IRI: https://w3id.org/brainteaser/ontology/schema/menstrualCycle
IRI: https://w3id.org/brainteaser/ontology/schema/observedResult
IRI: https://w3id.org/brainteaser/ontology/schema/onGene
IRI: https://w3id.org/brainteaser/ontology/schema/ownsSensingDevice
IRI: https://w3id.org/brainteaser/ontology/schema/performedTreatment
IRI: https://w3id.org/brainteaser/ontology/schema/pharmacologicSubstance
IRI: http://www.wurvoc.org/vocabularies/om-1.8/phenomenon
IRI: https://w3id.org/brainteaser/ontology/schema/placeLocation
IRI: https://w3id.org/brainteaser/ontology/schema/pregnancy
IRI: https://w3id.org/brainteaser/ontology/schema/relapse
IRI: https://w3id.org/brainteaser/ontology/schema/requiredMRI
IRI: https://w3id.org/brainteaser/ontology/schema/residence
IRI: https://w3id.org/brainteaser/ontology/schema/surgicalType
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/tested
IRI: https://w3id.org/brainteaser/ontology/schema/testedForOligoclonalBands
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/therapyType
IRI: https://w3id.org/brainteaser/ontology/schema/trauma
IRI: https://w3id.org/brainteaser/ontology/schema/undergo
IRI: http://www.wurvoc.org/vocabularies/om-1.8/unit_of_measure
IRI: http://www.wurvoc.org/vocabularies/om-1.8/value
IRI: https://w3id.org/brainteaser/ontology/schema/abdominalSurgery
IRI: https://w3id.org/brainteaser/ontology/schema/activeKilocalories
IRI: https://w3id.org/brainteaser/ontology/schema/activeTime
IRI: https://w3id.org/brainteaser/ontology/schema/activityDistance
IRI: https://w3id.org/brainteaser/ontology/schema/activityEndYear
IRI: https://w3id.org/brainteaser/ontology/schema/activityIntensity
IRI: https://w3id.org/brainteaser/ontology/schema/activityStartYear
IRI: https://w3id.org/brainteaser/ontology/schema/activitySteps
IRI: https://w3id.org/brainteaser/ontology/schema/activityStepsGoal
IRI: https://w3id.org/brainteaser/ontology/schema/activityStressDuration
IRI: https://w3id.org/brainteaser/ontology/schema/activityType
IRI: https://w3id.org/brainteaser/ontology/schema/adminstrationDose
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/administrationFrequency
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/administrationRoute
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/administrationUoM
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/ageBaseline
IRI: https://w3id.org/brainteaser/ontology/schema/ageDiagnosis
IRI: https://w3id.org/brainteaser/ontology/schema/ageOnset
IRI: https://w3id.org/brainteaser/ontology/schema/albuminCSF
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/alcoholUse
IRI: https://w3id.org/brainteaser/ontology/schema/alive
IRI: https://w3id.org/brainteaser/ontology/schema/alsfrs1
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/alsfrs10
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/alsfrs11
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/alsfrs12
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/alsfrs2
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/alsfrs3
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/alsfrs4
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/alsfrs5
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/alsfrs6
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/alsfrs7
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/alsfrs8
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/alsfrs9
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRS_R_WHO_COMPLETED
IRI: https://w3id.org/brainteaser/ontology/schema/alsfrsOriginal10
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q1
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q10
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q10alt
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q11
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q11alt
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q12
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q12alt
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q1alt
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q2
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q2alt
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q3
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q3alt
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q4
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q4alt
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q5
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q5A
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q5Aalt
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q5B
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q5Balt
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q6
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q6alt
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q7
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q7alt
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q8
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q8alt
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q9
IRI: https://w3id.org/brainteaser/ontology/schema/ALSFRSR_Q9alt
IRI: https://w3id.org/brainteaser/ontology/schema/alterationCSF
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/ambulationEDSS
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/ambulationEDSSAssistance
IRI: https://w3id.org/brainteaser/ontology/schema/ambulationEDSSObservedTime
IRI: https://w3id.org/brainteaser/ontology/schema/ambulationEDSSPatientDistance
IRI: https://w3id.org/brainteaser/ontology/schema/ambulationEDSSPatientTime
IRI: https://w3id.org/brainteaser/ontology/schema/ankleFootOrthosis
IRI: https://w3id.org/brainteaser/ontology/schema/ashworthLeftArm
IRI: https://w3id.org/brainteaser/ontology/schema/ashworthLeftLeg
IRI: https://w3id.org/brainteaser/ontology/schema/ashworthRightArm
IRI: https://w3id.org/brainteaser/ontology/schema/ashworthRightLeg
IRI: https://w3id.org/brainteaser/ontology/schema/autoimmuneDisease
IRI: https://w3id.org/brainteaser/ontology/schema/averageHeartRate
IRI: https://w3id.org/brainteaser/ontology/schema/averageMeasurement
IRI: https://w3id.org/brainteaser/ontology/schema/averageStressLevel
IRI: https://w3id.org/brainteaser/ontology/schema/awajiClinicalLMNBulbar
IRI: https://w3id.org/brainteaser/ontology/schema/awajiClinicalLMNCervical
IRI: https://w3id.org/brainteaser/ontology/schema/awajiClinicalLMNLumboSacral
IRI: https://w3id.org/brainteaser/ontology/schema/awajiClinicalLMNThoracic
IRI: https://w3id.org/brainteaser/ontology/schema/awajiClinicalUMNBulbar
IRI: https://w3id.org/brainteaser/ontology/schema/awajiClinicalUMNCervical
IRI: https://w3id.org/brainteaser/ontology/schema/awajiClinicalUMNLumboSacral
IRI: https://w3id.org/brainteaser/ontology/schema/awajiDiagnosisCriteria
IRI: https://w3id.org/brainteaser/ontology/schema/awajiEMGLMNBulbar
IRI: https://w3id.org/brainteaser/ontology/schema/awajiEMGLMNCervical
IRI: https://w3id.org/brainteaser/ontology/schema/awajiEMGLMNLumboSacral
IRI: https://w3id.org/brainteaser/ontology/schema/awajiEMGLMNThoracic
IRI: https://w3id.org/brainteaser/ontology/schema/awakeDuration
IRI: https://w3id.org/brainteaser/ontology/schema/axialOnset
IRI: https://w3id.org/brainteaser/ontology/schema/basalKilocalories
IRI: https://w3id.org/brainteaser/ontology/schema/baselineHeartRate
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatAI
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatC2a
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatC2d
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatCa
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatCd
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatCSI
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatCSIModified
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatCVI
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatCVSD
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatGI
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatHcvNN
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatHTI
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatIALS
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatIqrNN
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatMadNN
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatMeanNN
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatMedianNN
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatPAS
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatPI
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatPIP
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatPNN20
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatPNN50
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatPSS
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatRmssd
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatSD1
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatSD1a
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatSD1d
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatSD1SD2
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatSD2
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatSDANN1
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatSDANN2
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatSDANN5
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatSDNN
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatSDNNa
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatSDNNd
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatSDNNI
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatSDNNI1
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatSDNNI5
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatSDSD
IRI: https://w3id.org/brainteaser/ontology/schema/beatToBeatSI
IRI: https://w3id.org/brainteaser/ontology/schema/before20Years
IRI: https://w3id.org/brainteaser/ontology/schema/birthYear
IRI: https://w3id.org/brainteaser/ontology/schema/bmrKilocalories
IRI: https://w3id.org/brainteaser/ontology/schema/bmvtrTotal
IRI: https://w3id.org/brainteaser/ontology/schema/bodyMassIndex
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/bowelAndBladderEDSS
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/bowelAndBladderRelapse
IRI: https://w3id.org/brainteaser/ontology/schema/brainAtrophyMRI
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/brainstemEDSS
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/brainsetmRelapse
IRI: https://w3id.org/brainteaser/ontology/schema/bulbarOnset
IRI: https://w3id.org/brainteaser/ontology/schema/bulbarSubscore
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/burdenT1Lesions
IRI: https://w3id.org/brainteaser/ontology/schema/burdenT2Lesions
IRI: https://w3id.org/brainteaser/ontology/schema/bvmtrTrial1
IRI: https://w3id.org/brainteaser/ontology/schema/bvmtrTrial2
IRI: https://w3id.org/brainteaser/ontology/schema/bvmtrTrial3
IRI: https://w3id.org/brainteaser/ontology/schema/cardiacDisease
IRI: https://w3id.org/brainteaser/ontology/schema/cerebellarEDSS
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/cerebellarRelapse
IRI: https://w3id.org/brainteaser/ontology/schema/cerebralEDSS
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/cervicalSpineSurgery
IRI: https://w3id.org/brainteaser/ontology/schema/cervicalTrauma
IRI: https://w3id.org/brainteaser/ontology/schema/clinicallyEvaluatedEDSS
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/clinicalTrialDescription
IRI: https://w3id.org/brainteaser/ontology/schema/cognitivePhenotype
IRI: https://w3id.org/brainteaser/ontology/schema/cognitiveRelapse
IRI: https://w3id.org/brainteaser/ontology/schema/comorbiditySeverity
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/comorbidityTreatment
IRI: https://w3id.org/brainteaser/ontology/schema/componentName
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/concentrationMeasurement
IRI: https://w3id.org/brainteaser/ontology/schema/conditionEnd
IRI: https://w3id.org/brainteaser/ontology/schema/conditionStart
IRI: https://w3id.org/brainteaser/ontology/schema/contiguousLesionsMRI
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/cvltIITotalCorrect
IRI: https://w3id.org/brainteaser/ontology/schema/cvltIITotalInterferences
IRI: https://w3id.org/brainteaser/ontology/schema/cvltIITotalPerseverations
IRI: https://w3id.org/brainteaser/ontology/schema/cvltIITrial1Correct
IRI: https://w3id.org/brainteaser/ontology/schema/cvltIITrial1Interferences
IRI: https://w3id.org/brainteaser/ontology/schema/cvltIITrial1Perseverations
IRI: https://w3id.org/brainteaser/ontology/schema/cvltIITrial2Correct
IRI: https://w3id.org/brainteaser/ontology/schema/cvltIITrial2Interferences
IRI: https://w3id.org/brainteaser/ontology/schema/cvltIITrial2Perseverations
IRI: https://w3id.org/brainteaser/ontology/schema/cvltIITrial3Correct
IRI: https://w3id.org/brainteaser/ontology/schema/cvltIITrial3Interferences
IRI: https://w3id.org/brainteaser/ontology/schema/cvltIITrial3Perseverations
IRI: https://w3id.org/brainteaser/ontology/schema/cvltIITrial4Correct
IRI: https://w3id.org/brainteaser/ontology/schema/cvltIITrial4Interferences
IRI: https://w3id.org/brainteaser/ontology/schema/cvltIITrial4Perseverations
IRI: https://w3id.org/brainteaser/ontology/schema/cvltIITrial5Correct
IRI: https://w3id.org/brainteaser/ontology/schema/cvltIITrial5Interferences
IRI: https://w3id.org/brainteaser/ontology/schema/cvltIITrial5Perseverations
IRI: https://w3id.org/brainteaser/ontology/schema/dailyCigarettes
IRI: https://w3id.org/brainteaser/ontology/schema/deathCause
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/deathDate
IRI: https://w3id.org/brainteaser/ontology/schema/deepSleepDuration
IRI: https://w3id.org/brainteaser/ontology/schema/dextricity
IRI: https://w3id.org/brainteaser/ontology/schema/diabetes
IRI: https://w3id.org/brainteaser/ontology/schema/diagnosticCriteria
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/diagnosticCriteriaLabel
IRI: https://w3id.org/brainteaser/ontology/schema/diagnosticDelay
IRI: https://w3id.org/brainteaser/ontology/schema/diet
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/dominantHandTrial1
IRI: https://w3id.org/brainteaser/ontology/schema/dominantHandTrial2
IRI: https://w3id.org/brainteaser/ontology/schema/dominantHandTrialDone
IRI: https://w3id.org/brainteaser/ontology/schema/DYMUS_Q1
IRI: https://w3id.org/brainteaser/ontology/schema/DYMUS_Q10
IRI: https://w3id.org/brainteaser/ontology/schema/DYMUS_Q2
IRI: https://w3id.org/brainteaser/ontology/schema/DYMUS_Q3
IRI: https://w3id.org/brainteaser/ontology/schema/DYMUS_Q4
IRI: https://w3id.org/brainteaser/ontology/schema/DYMUS_Q5
IRI: https://w3id.org/brainteaser/ontology/schema/DYMUS_Q6
IRI: https://w3id.org/brainteaser/ontology/schema/DYMUS_Q7
IRI: https://w3id.org/brainteaser/ontology/schema/DYMUS_Q8
IRI: https://w3id.org/brainteaser/ontology/schema/DYMUS_Q9
IRI: https://w3id.org/brainteaser/ontology/schema/dyslipidemia
IRI: https://w3id.org/brainteaser/ontology/schema/ecasALSnonSpecificScore
IRI: https://w3id.org/brainteaser/ontology/schema/ecasALSspecificScore
IRI: https://w3id.org/brainteaser/ontology/schema/ecasCarerInterviewScore
IRI: https://w3id.org/brainteaser/ontology/schema/ecasExecutiveAlternation
IRI: https://w3id.org/brainteaser/ontology/schema/ecasExecutiveFluencyS
IRI: https://w3id.org/brainteaser/ontology/schema/ecasExecutiveFluencyT
IRI: https://w3id.org/brainteaser/ontology/schema/ecasExecutiveScore
IRI: https://w3id.org/brainteaser/ontology/schema/ecasExecutiveSentenceCompletion
IRI: https://w3id.org/brainteaser/ontology/schema/ecasHasCarerInterview
IRI: https://w3id.org/brainteaser/ontology/schema/ecasLanguageComprehension
IRI: https://w3id.org/brainteaser/ontology/schema/ecasLanguageNaming
IRI: https://w3id.org/brainteaser/ontology/schema/ecasLanguageScore
IRI: https://w3id.org/brainteaser/ontology/schema/ecasLanguageSpelling
IRI: https://w3id.org/brainteaser/ontology/schema/ecasMemoryDelayedRecall
IRI: https://w3id.org/brainteaser/ontology/schema/ecasMemoryDelayedRecognition
IRI: https://w3id.org/brainteaser/ontology/schema/ecasMemoryImmediateRecall
IRI: https://w3id.org/brainteaser/ontology/schema/ecasMemoryScore
IRI: https://w3id.org/brainteaser/ontology/schema/ecasPsychosisScreenScore
IRI: https://w3id.org/brainteaser/ontology/schema/ecasReverseDigitSpan
IRI: https://w3id.org/brainteaser/ontology/schema/ecasSocialCognition
IRI: https://w3id.org/brainteaser/ontology/schema/ecasSymptomsApathyOrInertia
IRI: https://w3id.org/brainteaser/ontology/schema/ecasSymptomsChangePrevBehaviour
IRI: https://w3id.org/brainteaser/ontology/schema/ecasSymptomsDisinhibition
IRI: https://w3id.org/brainteaser/ontology/schema/ecasSymptomsHyperorality
IRI: https://w3id.org/brainteaser/ontology/schema/ecasSymptomsLossSympathyOrEmpathy
IRI: https://w3id.org/brainteaser/ontology/schema/ecasSymptomsPerseverativeSterotyped
IRI: https://w3id.org/brainteaser/ontology/schema/ecasTotalScore
IRI: https://w3id.org/brainteaser/ontology/schema/ecasVerbalFluencyScore
IRI: https://w3id.org/brainteaser/ontology/schema/ecasVisuospatialCubeCounting
IRI: https://w3id.org/brainteaser/ontology/schema/ecasVisuospatialDotCounting
IRI: https://w3id.org/brainteaser/ontology/schema/ecasVisuospatialNumberLocation
IRI: https://w3id.org/brainteaser/ontology/schema/ecasVisuospatialScore
IRI: https://w3id.org/brainteaser/ontology/schema/educationLevel
IRI: https://w3id.org/brainteaser/ontology/schema/endReason
IRI: https://w3id.org/brainteaser/ontology/schema/endTherapyReason
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/EQ5D5L_Q1
IRI: https://w3id.org/brainteaser/ontology/schema/EQ5D5L_Q2
IRI: https://w3id.org/brainteaser/ontology/schema/EQ5D5L_Q3
IRI: https://w3id.org/brainteaser/ontology/schema/EQ5D5L_Q4
IRI: https://w3id.org/brainteaser/ontology/schema/EQ5D5L_Q5
IRI: https://w3id.org/brainteaser/ontology/schema/EQ5D5L_Q6
IRI: https://w3id.org/brainteaser/ontology/schema/eventEnd
IRI: https://w3id.org/brainteaser/ontology/schema/eventStart
IRI: https://w3id.org/brainteaser/ontology/schema/everSmoked
IRI: https://w3id.org/brainteaser/ontology/schema/evokedPotentialsLocation
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/evokedPotentialsResponse
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/evokedPotentialsType
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/factorsAffectedHandTrial
IRI: https://w3id.org/brainteaser/ontology/schema/familiarHistoryALS
IRI: https://w3id.org/brainteaser/ontology/schema/floorsClimbed
IRI: https://w3id.org/brainteaser/ontology/schema/floorsClimbedGoal
IRI: https://w3id.org/brainteaser/ontology/schema/gadoliniumLesionsT1MRI
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/gadoliniumLesionsT1MRINumber
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/generalizedOnset
IRI: https://w3id.org/brainteaser/ontology/schema/genomeSequencing
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/glucoseCSF
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/goldCoastExcludingOthers
IRI: https://w3id.org/brainteaser/ontology/schema/goldCoastProgressiveMotorImpairment
IRI: https://w3id.org/brainteaser/ontology/schema/goldCoastUMNandLMN
IRI: https://w3id.org/brainteaser/ontology/schema/granulocytesCSF
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/hadMajorTrauma
IRI: https://w3id.org/brainteaser/ontology/schema/HADS_Q1
IRI: https://w3id.org/brainteaser/ontology/schema/HADS_Q10
IRI: https://w3id.org/brainteaser/ontology/schema/HADS_Q11
IRI: https://w3id.org/brainteaser/ontology/schema/HADS_Q12
IRI: https://w3id.org/brainteaser/ontology/schema/HADS_Q13
IRI: https://w3id.org/brainteaser/ontology/schema/HADS_Q14
IRI: https://w3id.org/brainteaser/ontology/schema/HADS_Q2
IRI: https://w3id.org/brainteaser/ontology/schema/HADS_Q3
IRI: https://w3id.org/brainteaser/ontology/schema/HADS_Q4
IRI: https://w3id.org/brainteaser/ontology/schema/HADS_Q5
IRI: https://w3id.org/brainteaser/ontology/schema/HADS_Q6
IRI: https://w3id.org/brainteaser/ontology/schema/HADS_Q7
IRI: https://w3id.org/brainteaser/ontology/schema/HADS_Q8
IRI: https://w3id.org/brainteaser/ontology/schema/HADS_Q9
IRI: https://w3id.org/brainteaser/ontology/schema/hadSurgicalIntervention
IRI: https://w3id.org/brainteaser/ontology/schema/hasMenstrualCycle
IRI: https://w3id.org/brainteaser/ontology/schema/hasMutation
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/headNeckSurgery
IRI: https://w3id.org/brainteaser/ontology/schema/headTrauma
IRI: https://w3id.org/brainteaser/ontology/schema/height
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/highStressDuration
IRI: https://w3id.org/brainteaser/ontology/schema/homeWalkingStatus
IRI: https://w3id.org/brainteaser/ontology/schema/hypertension
IRI: https://w3id.org/brainteaser/ontology/schema/IgGIndexCSF
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/immunoglobulinGCSF
has characteristics: functional
IRI: http://www.w3.org/2006/time#inXSDDateTime
IRI: https://w3id.org/brainteaser/ontology/schema/intensityDurationGoal
IRI: https://w3id.org/brainteaser/ontology/schema/isMenstrualCycleRegular
IRI: https://w3id.org/brainteaser/ontology/schema/kingsValue
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/lesionsT1MRI
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/lesionsT2MRI
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/leukocytesCountCSF
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/lightSleepDuration
IRI: https://w3id.org/brainteaser/ontology/schema/limbsOnset
IRI: https://w3id.org/brainteaser/ontology/schema/linearTrendHeartRate
IRI: https://w3id.org/brainteaser/ontology/schema/lowerLimbSurgery
IRI: https://w3id.org/brainteaser/ontology/schema/lowStressDuration
IRI: https://w3id.org/brainteaser/ontology/schema/lumboSacralSpineSurgery
IRI: https://w3id.org/brainteaser/ontology/schema/lumboSacralTrauma
IRI: https://w3id.org/brainteaser/ontology/schema/lymphocytesCSF
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/maritalStatus
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/maxHeartRate
IRI: https://w3id.org/brainteaser/ontology/schema/maxStressLevel
IRI: https://w3id.org/brainteaser/ontology/schema/maxTimeHeartRate
IRI: https://w3id.org/brainteaser/ontology/schema/measuredLevel
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/measurementCalibrated
IRI: https://w3id.org/brainteaser/ontology/schema/mediumStressDuration
IRI: https://w3id.org/brainteaser/ontology/schema/menopauseStatus
IRI: https://w3id.org/brainteaser/ontology/schema/metersWalkedFirstMin
IRI: https://w3id.org/brainteaser/ontology/schema/metersWalkedTwoMins
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q1
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q10
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q11
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q12
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q13
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q14
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q15
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q16
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q17
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q18
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q19
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q2
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q20
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q21
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q3
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q4
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q5
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q6
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q7
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q8
IRI: https://w3id.org/brainteaser/ontology/schema/MFIS_Q9
IRI: https://w3id.org/brainteaser/ontology/schema/minHeartRate
IRI: https://w3id.org/brainteaser/ontology/schema/minTimeHeartRate
IRI: https://w3id.org/brainteaser/ontology/schema/mitosValue
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/moderateIntensityDuration
IRI: https://w3id.org/brainteaser/ontology/schema/moreThan10PercentWeightloss
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/motorSubscore
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/MSInPaediatricAge
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q1
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q10
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q11
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q12
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q13
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q14
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q15
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q16
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q17
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q18
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q19
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q2
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q20
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q21
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q22
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q23
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q24
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q25
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q26
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q27
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q28
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q29
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q3
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q30
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q31
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q32
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q33
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q34
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q35
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q36
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q37
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q38
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q39
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q4
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q40
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q41
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q42
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q43
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q44
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q45
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q46
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q47
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q48
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q49
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q5
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q50
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q51
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q52
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q53
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q54
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q6
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q7
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q8
IRI: https://w3id.org/brainteaser/ontology/schema/MSQOL54_Q9
IRI: https://w3id.org/brainteaser/ontology/schema/multipleSclerosisType
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/mutationType
IRI: https://w3id.org/brainteaser/ontology/schema/neckTrauma
IRI: https://w3id.org/brainteaser/ontology/schema/newOrEnlargedLesionsT2MRI
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/newOrEnlargedLesionsT2MRINumber
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/nonDominantHandTrial1
IRI: https://w3id.org/brainteaser/ontology/schema/nonDominantHandTrial2
IRI: https://w3id.org/brainteaser/ontology/schema/nonDominantHandTrialDone
IRI: https://w3id.org/brainteaser/ontology/schema/normalLevel
has characteristics: functional
IRI: http://www.w3.org/2004/02/skos/core#notation
IRI: https://w3id.org/brainteaser/ontology/schema/notes
IRI: https://w3id.org/brainteaser/ontology/schema/numberSpinalCordLesions
IRI: http://www.wurvoc.org/vocabularies/om-1.8/numerical_value
IRI: https://w3id.org/brainteaser/ontology/schema/oligoclonalBandsNumber
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/oligoclonalBandsPresence
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/oligoclonalBandsType
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/onsetElapsedTime
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/originalALSFRS
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/otherFactorsAffectedHandTrial
IRI: https://w3id.org/brainteaser/ontology/schema/otherOnset
IRI: https://w3id.org/brainteaser/ontology/schema/otherRelapse
IRI: https://w3id.org/brainteaser/ontology/schema/outsideWalkingStatus
IRI: https://w3id.org/brainteaser/ontology/schema/packYear
IRI: https://w3id.org/brainteaser/ontology/schema/participationEnd
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/participationStart
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/pathologicalJCCSF
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/pelvicSurgery
IRI: https://w3id.org/brainteaser/ontology/schema/PFTs_FEV1_L
IRI: https://w3id.org/brainteaser/ontology/schema/PFTs_FEV1_P
IRI: https://w3id.org/brainteaser/ontology/schema/PFTs_FEV6_L
IRI: https://w3id.org/brainteaser/ontology/schema/PFTs_FEV6_P
IRI: https://w3id.org/brainteaser/ontology/schema/phenotype
IRI: https://w3id.org/brainteaser/ontology/schema/placeArea
IRI: https://w3id.org/brainteaser/ontology/schema/positivityResult
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/predominantLimbsImpairment
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/predominantLimbsSide
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/predominantMotorNeuron
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/pregnancyComplications
IRI: https://w3id.org/brainteaser/ontology/schema/pregnancyEnd
IRI: https://w3id.org/brainteaser/ontology/schema/pregnancyEndEvent
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/pregnancyStart
IRI: https://w3id.org/brainteaser/ontology/schema/primaryNeoplasm
IRI: https://w3id.org/brainteaser/ontology/schema/privacyPreservingDate
IRI: https://w3id.org/brainteaser/ontology/schema/privacyPreservingDeathDate
IRI: https://w3id.org/brainteaser/ontology/schema/procedureEnd
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/procedureStart
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/proteinsCSF
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q1
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q10
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q11A
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q11B
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q11C
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q11D
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q11E
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q2
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q3
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q4
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q5A
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q5B
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q5C
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q5D
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q5E
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q5F
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q5G
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q5H
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q5I
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q5J_1
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q5J_2
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q6
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q7
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q8
IRI: https://w3id.org/brainteaser/ontology/schema/PSQI_Q9
IRI: https://w3id.org/brainteaser/ontology/schema/pulmonaryAOPAbs
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/pulmonaryFEV6
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/pulmonaryFVCLyingDown
IRI: https://w3id.org/brainteaser/ontology/schema/pulmonaryFVCLyingDownPercentage
IRI: https://w3id.org/brainteaser/ontology/schema/pulmonaryFVCRel
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/pulmonaryFVCSitting
IRI: https://w3id.org/brainteaser/ontology/schema/pulmonaryFVCSittingPercentage
IRI: https://w3id.org/brainteaser/ontology/schema/pulmonaryMEP
IRI: https://w3id.org/brainteaser/ontology/schema/pulmonaryMEPAbs
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/pulmonaryMEPpercentage
IRI: https://w3id.org/brainteaser/ontology/schema/pulmonaryMIP
IRI: https://w3id.org/brainteaser/ontology/schema/pulmonaryMIPAbs
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/pulmonaryMIPpercentage
IRI: https://w3id.org/brainteaser/ontology/schema/pulmonaryNIP
IRI: https://w3id.org/brainteaser/ontology/schema/pulmonaryPCF
IRI: https://w3id.org/brainteaser/ontology/schema/pulmonaryPCO2
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/pulmonaryPO2
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/pulmonaryVCAbs
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/pyramidalEDSS
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/pyramidalRelapse
IRI: https://w3id.org/brainteaser/ontology/schema/quadraticTrendHeartRate
IRI: https://w3id.org/brainteaser/ontology/schema/questionnaireSubmitted
IRI: https://w3id.org/brainteaser/ontology/schema/quotientAlbuminCSF
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/r2HeartRate
IRI: https://w3id.org/brainteaser/ontology/schema/reasonIrregularCycle
IRI: https://w3id.org/brainteaser/ontology/schema/reasonNoMenstrualCycle
IRI: https://w3id.org/brainteaser/ontology/schema/reasonWalkTestNotCompleted
IRI: https://w3id.org/brainteaser/ontology/schema/relapseClinicalTherapy
IRI: https://w3id.org/brainteaser/ontology/schema/relapseCortisoneTherapy
IRI: https://w3id.org/brainteaser/ontology/schema/relapseDuration
IRI: https://w3id.org/brainteaser/ontology/schema/relapseEnd
IRI: https://w3id.org/brainteaser/ontology/schema/relapseHospitalAdmission
IRI: https://w3id.org/brainteaser/ontology/schema/relapseImpactOnADL
IRI: https://w3id.org/brainteaser/ontology/schema/relapseRecovery
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/relapseSequela
IRI: https://w3id.org/brainteaser/ontology/schema/relapseSeverity
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/relapseStart
IRI: https://w3id.org/brainteaser/ontology/schema/relativeEndDate
IRI: https://w3id.org/brainteaser/ontology/schema/relativeStartDate
IRI: https://w3id.org/brainteaser/ontology/schema/remSleep
IRI: https://w3id.org/brainteaser/ontology/schema/repirationSampEn
IRI: https://w3id.org/brainteaser/ontology/schema/residenceEnd
IRI: https://w3id.org/brainteaser/ontology/schema/residenceStart
IRI: https://w3id.org/brainteaser/ontology/schema/respirationAlpha1DimMean
IRI: https://w3id.org/brainteaser/ontology/schema/respirationAlpha1DimRange
IRI: https://w3id.org/brainteaser/ontology/schema/respirationAlpha1ExpMean
IRI: https://w3id.org/brainteaser/ontology/schema/respirationAlpha1ExpRange
IRI: https://w3id.org/brainteaser/ontology/schema/respirationAlpha2DimMean
IRI: https://w3id.org/brainteaser/ontology/schema/respirationAlpha2DimRange
IRI: https://w3id.org/brainteaser/ontology/schema/respirationAlpha2ExpMean
IRI: https://w3id.org/brainteaser/ontology/schema/respirationAlpha2ExpRange
IRI: https://w3id.org/brainteaser/ontology/schema/respirationApEn
IRI: https://w3id.org/brainteaser/ontology/schema/respirationDFAalpha1
IRI: https://w3id.org/brainteaser/ontology/schema/respirationDFAalpha2
IRI: https://w3id.org/brainteaser/ontology/schema/respirationRMSSD
IRI: https://w3id.org/brainteaser/ontology/schema/respirationSD1
IRI: https://w3id.org/brainteaser/ontology/schema/respirationSD2
IRI: https://w3id.org/brainteaser/ontology/schema/respirationSD2SD1
IRI: https://w3id.org/brainteaser/ontology/schema/respirationSDBB
IRI: https://w3id.org/brainteaser/ontology/schema/respirationSDSD
IRI: https://w3id.org/brainteaser/ontology/schema/respiratorySubscore
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/restingHeartRate
IRI: https://w3id.org/brainteaser/ontology/schema/restStressDuration
IRI: https://w3id.org/brainteaser/ontology/schema/retiredAtDiagnosis
IRI: https://w3id.org/brainteaser/ontology/schema/revisedALSFRS
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/sdmtCorrect
IRI: https://w3id.org/brainteaser/ontology/schema/sdmtIncorrect
IRI: https://w3id.org/brainteaser/ontology/schema/sensoryEDSS
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/sensoryRelapse
IRI: https://w3id.org/brainteaser/ontology/schema/sex
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/sexuality
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/SLEEP_ALS_Q1
IRI: https://w3id.org/brainteaser/ontology/schema/slope
IRI: https://w3id.org/brainteaser/ontology/schema/smokersNumber
IRI: https://w3id.org/brainteaser/ontology/schema/smokingExpositionType
IRI: https://w3id.org/brainteaser/ontology/schema/spinalCordLesions
IRI: https://w3id.org/brainteaser/ontology/schema/spo2AOD100
IRI: https://w3id.org/brainteaser/ontology/schema/spo2AODMax
IRI: https://w3id.org/brainteaser/ontology/schema/spo2AV
IRI: https://w3id.org/brainteaser/ontology/schema/spo2CA
IRI: https://w3id.org/brainteaser/ontology/schema/spo2CT
IRI: https://w3id.org/brainteaser/ontology/schema/spo2DI
IRI: https://w3id.org/brainteaser/ontology/schema/spo2M
IRI: https://w3id.org/brainteaser/ontology/schema/spo2MED
IRI: https://w3id.org/brainteaser/ontology/schema/spo2Min
IRI: https://w3id.org/brainteaser/ontology/schema/spo2ODI
IRI: https://w3id.org/brainteaser/ontology/schema/spo2P
IRI: https://w3id.org/brainteaser/ontology/schema/spo2POD
IRI: https://w3id.org/brainteaser/ontology/schema/spo2RG
IRI: https://w3id.org/brainteaser/ontology/schema/spo2SD
IRI: https://w3id.org/brainteaser/ontology/schema/spo2ZC
IRI: https://w3id.org/brainteaser/ontology/schema/stdHeartRate
IRI: https://w3id.org/brainteaser/ontology/schema/steps12am6am
IRI: https://w3id.org/brainteaser/ontology/schema/steps12pm6pm
IRI: https://w3id.org/brainteaser/ontology/schema/steps6am12pm
IRI: https://w3id.org/brainteaser/ontology/schema/stressDuration
IRI: https://w3id.org/brainteaser/ontology/schema/stressQualifier
IRI: https://w3id.org/brainteaser/ontology/schema/stroke
IRI: https://w3id.org/brainteaser/ontology/schema/sunExposure
IRI: https://w3id.org/brainteaser/ontology/schema/symptomClinicalName
IRI: https://w3id.org/brainteaser/ontology/schema/symptomEnd
IRI: https://w3id.org/brainteaser/ontology/schema/symptomGrade
IRI: https://w3id.org/brainteaser/ontology/schema/symptomStart
IRI: https://w3id.org/brainteaser/ontology/schema/symptomType
IRI: https://w3id.org/brainteaser/ontology/schema/symptomValue
IRI: https://w3id.org/brainteaser/ontology/schema/thoracicSpineSurgery
IRI: https://w3id.org/brainteaser/ontology/schema/thoracicSurgery
IRI: https://w3id.org/brainteaser/ontology/schema/thoracicTrauma
IRI: https://w3id.org/brainteaser/ontology/schema/thyroidDisorder
IRI: https://w3id.org/brainteaser/ontology/schema/totalCalories
IRI: https://w3id.org/brainteaser/ontology/schema/totalLesionsT2MRI
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/totalSleepDuration
IRI: https://w3id.org/brainteaser/ontology/schema/traumaDate
IRI: https://w3id.org/brainteaser/ontology/schema/traumaDescription
IRI: https://w3id.org/brainteaser/ontology/schema/unitOfMeasure
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/unmeasurableSleepDuration
IRI: https://w3id.org/brainteaser/ontology/schema/upperLimbSurgery
IRI: https://w3id.org/brainteaser/ontology/schema/validMeasurements
IRI: https://w3id.org/brainteaser/ontology/schema/VASQ1
IRI: https://w3id.org/brainteaser/ontology/schema/vigorousIntensityDuration
IRI: https://w3id.org/brainteaser/ontology/schema/virusJCTestedCSF
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/visualAcuityLeft
IRI: https://w3id.org/brainteaser/ontology/schema/visualAcuityRight
IRI: https://w3id.org/brainteaser/ontology/schema/visualEDSS
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/visualEvokedPotentialAmplitude
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/visualEvokedPotentialLatency
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/visualRelapse
IRI: https://w3id.org/brainteaser/ontology/schema/walkAssistiveDevice
IRI: https://w3id.org/brainteaser/ontology/schema/walkAssistiveDeviceUsed
IRI: https://w3id.org/brainteaser/ontology/schema/walkingTime10Meters
IRI: https://w3id.org/brainteaser/ontology/schema/walkingTime25Steps
IRI: https://w3id.org/brainteaser/ontology/schema/walkTestNotCompleted
IRI: https://w3id.org/brainteaser/ontology/schema/wdmDuration
IRI: https://w3id.org/brainteaser/ontology/schema/wdmStartTime
IRI: https://w3id.org/brainteaser/ontology/schema/weeklyFrequency
IRI: https://w3id.org/brainteaser/ontology/schema/weight
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/WEIGHT_Q1
IRI: https://w3id.org/brainteaser/ontology/schema/whoThresholdExceeded
IRI: https://w3id.org/brainteaser/ontology/schema/zipCode
has characteristics: functional
IRI: https://w3id.org/brainteaser/ontology/schema/zipHashed
has characteristics: functional
IRI: http://purl.obolibrary.org/obo/NCIT_C12664
IRI: http://purl.obolibrary.org/obo/NCIT_C26682
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/177250006
IRI: http://purl.bioontology.org/ontology/ATC/C09AA
IRI: http://purl.bioontology.org/ontology/ATC/N06BX12
IRI: http://purl.bioontology.org/ontology/ATC/N02BA01
IRI: http://purl.bioontology.org/ontology/ATC/N02BA51
IRI: http://purl.bioontology.org/ontology/ATC/J05AB01
IRI: http://purl.obolibrary.org/obo/NCIT_C129644
IRI: http://purl.obolibrary.org/obo/NCIT_C2851
IRI: http://purl.obolibrary.org/obo/NCIT_C26934
IRI: http://purl.obolibrary.org/obo/NCIT_C3171
IRI: http://purl.obolibrary.org/obo/NCIT_C35204
IRI: http://purl.obolibrary.org/obo/NCIT_C123215
IRI: http://purl.obolibrary.org/obo/NCIT_C27043
IRI: http://purl.obolibrary.org/obo/NCIT_C97142
IRI: http://purl.bioontology.org/ontology/ATC/A16AA02
IRI: http://purl.obolibrary.org/obo/NCIT_C2852
IRI: http://purl.obolibrary.org/obo/OAE_0000220
IRI: http://data.europa.eu/esco/isco/C12
IRI: http://purl.obolibrary.org/obo/OAE_0000005
IRI: http://purl.bioontology.org/ontology/ATC/C09
IRI: http://purl.obolibrary.org/obo/NCIT_C116374
IRI: http://data.europa.eu/esco/isco/C92
IRI: http://purl.obolibrary.org/obo/NCIT_C26948
IRI: http://purl.obolibrary.org/obo/NCIT_C93040
IRI: http://purl.bioontology.org/ontology/ATC/L01ED03
IRI: http://purl.bioontology.org/ontology/ATC/L04AA34
IRI: http://purl.bioontology.org/ontology/ATC/G04CA01
IRI: http://purl.bioontology.org/ontology/ATC/A
IRI: http://purl.bioontology.org/ontology/ATC/V07
IRI: http://purl.bioontology.org/ontology/ATC/V03
IRI: http://purl.bioontology.org/ontology/ATC/V01
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/389145006
IRI: http://purl.obolibrary.org/obo/NCIT_C79532
IRI: http://purl.obolibrary.org/obo/NCIT_C61595
IRI: http://purl.bioontology.org/ontology/ATC/N05BA12
IRI: http://purl.obolibrary.org/obo/NCIT_C2866
IRI: http://purl.bioontology.org/ontology/ATC/N04BB01
IRI: http://purl.bioontology.org/ontology/ATC/C02KX02
IRI: http://purl.obolibrary.org/obo/NCIT_C61443
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/1144987003
IRI: http://purl.bioontology.org/ontology/ATC/C01BD01
IRI: http://purl.bioontology.org/ontology/ATC/N06AA09
IRI: http://purl.bioontology.org/ontology/ATC/N06CA01
IRI: http://purl.bioontology.org/ontology/ATC/C08CA01
IRI: http://www.wurvoc.org/vocabularies/om-1.8/amount_of_substance_of_a_system_that_contains_as_many_elementary_entities_as_there_are_atoms_in_0.012_kilogram_of_carbon_12
IRI: http://purl.bioontology.org/ontology/ATC/J01CR02
IRI: http://purl.obolibrary.org/obo/NCIT_C34373
IRI: http://purl.bioontology.org/ontology/ATC/A14
IRI: http://purl.bioontology.org/ontology/ATC/N02
IRI: http://purl.obolibrary.org/obo/NCIT_C2869
IRI: http://purl.bioontology.org/ontology/ATC/N01
IRI: http://purl.obolibrary.org/obo/NCIT_C112175
IRI: http://purl.obolibrary.org/obo/NCIT_C26989
IRI: http://purl.obolibrary.org/obo/NCIT_C8486
IRI: http://purl.obolibrary.org/obo/NCIT_C8668
IRI: http://purl.obolibrary.org/obo/NCIT_C60785
IRI: http://purl.obolibrary.org/obo/NCIT_C2875
IRI: http://purl.obolibrary.org/obo/NCIT_C116369
IRI: http://purl.bioontology.org/ontology/ATC/P02
IRI: http://purl.bioontology.org/ontology/ATC/D10
IRI: http://purl.bioontology.org/ontology/ATC/N04
IRI: http://purl.bioontology.org/ontology/ATC/B03
IRI: http://purl.bioontology.org/ontology/ATC/J01
IRI: http://purl.bioontology.org/ontology/ATC/D06
IRI: http://purl.obolibrary.org/obo/NCIT_C264
IRI: http://purl.bioontology.org/ontology/ATC/N06A
IRI: http://purl.bioontology.org/ontology/ATC/A07
IRI: http://purl.bioontology.org/ontology/ATC/A04
IRI: http://purl.bioontology.org/ontology/ATC/N03
IRI: http://purl.bioontology.org/ontology/ATC/D01
IRI: http://purl.bioontology.org/ontology/ATC/M04
IRI: http://purl.bioontology.org/ontology/ATC/B02
IRI: http://purl.bioontology.org/ontology/ATC/R06
IRI: http://purl.bioontology.org/ontology/ATC/C02
IRI: http://purl.bioontology.org/ontology/ATC/J
IRI: http://purl.bioontology.org/ontology/ATC/M01
IRI: http://purl.bioontology.org/ontology/ATC/M01A
IRI: http://purl.bioontology.org/ontology/ATC/J04
IRI: http://purl.bioontology.org/ontology/ATC/J02
IRI: http://purl.bioontology.org/ontology/ATC/L01
IRI: http://purl.bioontology.org/ontology/ATC/L
IRI: http://purl.bioontology.org/ontology/ATC/A08
IRI: http://purl.bioontology.org/ontology/ATC/P
IRI: http://purl.bioontology.org/ontology/ATC/P01
IRI: http://purl.bioontology.org/ontology/ATC/D04
IRI: http://purl.bioontology.org/ontology/ATC/D05
IRI: http://purl.bioontology.org/ontology/ATC/D08
IRI: http://purl.bioontology.org/ontology/ATC/B01
IRI: http://purl.bioontology.org/ontology/ATC/J05
IRI: http://purl.obolibrary.org/obo/NCIT_C2878
IRI: http://purl.bioontology.org/ontology/ATC/N05B
IRI: http://purl.bioontology.org/ontology/ATC/B01AF02
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/80146002
IRI: http://purl.bioontology.org/ontology/ATC/A15
IRI: http://purl.obolibrary.org/obo/NCIT_C28539
IRI: http://purl.bioontology.org/ontology/ATC/N05AX12
IRI: http://data.europa.eu/esco/isco/C0
IRI: http://data.europa.eu/esco/isco/C03
IRI: http://purl.obolibrary.org/obo/NCIT_C2881
IRI: http://purl.obolibrary.org/obo/NCIT_C2883
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/315280000
IRI: http://purl.obolibrary.org/obo/NCIT_C34932
IRI: http://data.europa.eu/esco/isco/C82
IRI: http://purl.obolibrary.org/obo/NCIT_C28397
IRI: http://purl.bioontology.org/ontology/ATC/C07AB03
IRI: http://purl.bioontology.org/ontology/ATC/C07CB03
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/24079001
IRI: http://purl.obolibrary.org/obo/NCIT_C50466
IRI: http://purl.obolibrary.org/obo/NCIT_C51224
IRI: http://purl.bioontology.org/ontology/ATC/C10AA05
IRI: http://purl.obolibrary.org/obo/NCIT_C96569
IRI: http://purl.obolibrary.org/obo/NCIT_C2889
IRI: http://purl.obolibrary.org/obo/NCIT_C99383
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/183005
IRI: http://purl.bioontology.org/ontology/ATC/L04AX01
IRI: http://purl.obolibrary.org/obo/NCIT_C3457
IRI: http://purl.bioontology.org/ontology/ATC/M03BX01
IRI: http://purl.obolibrary.org/obo/NCIT_C26704
IRI: http://purl.obolibrary.org/obo/NCIT_C34935
IRI: http://purl.bioontology.org/ontology/ATC/N03AA04
IRI: http://purl.bioontology.org/ontology/ATC/C08CA12
IRI: http://purl.obolibrary.org/obo/NCIT_C156767
IRI: http://purl.bioontology.org/ontology/ATC/L03AX03
IRI: https://w3id.org/brainteaser/ontology/schema/BehavioralImpairment
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/193093009
IRI: http://purl.obolibrary.org/obo/NCIT_C2897
IRI: http://purl.bioontology.org/ontology/ATC/C07
IRI: http://purl.bioontology.org/ontology/ATC/N07CA01
IRI: http://purl.bioontology.org/ontology/ATC/H02AB01
IRI: https://w3id.org/brainteaser/ontology/schema/N07XX09
IRI: http://purl.bioontology.org/ontology/ATC/A05
IRI: http://purl.bioontology.org/ontology/ATC/A05A
IRI: http://purl.bioontology.org/ontology/ATC/A11HA05
IRI: http://purl.obolibrary.org/obo/NCIT_C34423
IRI: http://purl.bioontology.org/ontology/ATC/C07AB07
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/315240009
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/18167009
IRI: http://purl.obolibrary.org/obo/NCIT_C112183
IRI: http://purl.obolibrary.org/obo/NCIT_C118723
IRI: http://purl.bioontology.org/ontology/ATC/B
IRI: http://purl.bioontology.org/ontology/ATC/B05
IRI: http://purl.obolibrary.org/obo/NCIT_C12431
IRI: http://purl.bioontology.org/ontology/ATC/M03AX01
IRI: http://purl.obolibrary.org/obo/NCIT_C9175
IRI: http://purl.obolibrary.org/obo/NCIT_C37920
IRI: http://purl.obolibrary.org/obo/NCIT_C12439
IRI: http://purl.obolibrary.org/obo/NCIT_C12441
IRI: http://purl.obolibrary.org/obo/NCIT_C4872
IRI: http://purl.obolibrary.org/obo/NCIT_C4017
IRI: http://purl.bioontology.org/ontology/ATC/N05BA08
IRI: http://purl.obolibrary.org/obo/NCIT_C111201
IRI: http://purl.bioontology.org/ontology/ATC/N05CD09
IRI: http://purl.obolibrary.org/obo/NCIT_C84601
IRI: http://data.europa.eu/esco/isco/C71
IRI: http://data.europa.eu/esco/isco/C33
IRI: http://data.europa.eu/esco/isco/C24
IRI: http://purl.obolibrary.org/obo/NCIT_C101398
IRI: http://purl.bioontology.org/ontology/ATC/N04BC06
IRI: http://purl.bioontology.org/ontology/ATC/A11CC06
IRI: http://purl.bioontology.org/ontology/ATC/C08
IRI: http://purl.bioontology.org/ontology/ATC/V03AF03
IRI: http://purl.bioontology.org/ontology/ATC/H05
IRI: http://purl.bioontology.org/ontology/ATC/A12AX
IRI: http://purl.bioontology.org/ontology/ATC/C09DA06
IRI: http://purl.bioontology.org/ontology/ATC/N03AX24
IRI: http://purl.bioontology.org/ontology/ATC/N02BG10
IRI: http://purl.bioontology.org/ontology/ATC/N03AF01
IRI: http://purl.obolibrary.org/obo/NCIT_C2916
IRI: http://purl.bioontology.org/ontology/ATC/C01
IRI: http://purl.obolibrary.org/obo/NCIT_C2931
IRI: http://purl.bioontology.org/ontology/ATC/C
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/250980009
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/14045001
IRI: http://purl.obolibrary.org/obo/NCIT_C3086
IRI: http://purl.bioontology.org/ontology/ATC/J01DD04
IRI: http://purl.obolibrary.org/obo/NCIT_C26714
IRI: http://purl.obolibrary.org/obo/NCIT_C4802
IRI: http://purl.obolibrary.org/obo/NCIT_C12445
IRI: http://purl.obolibrary.org/obo/NCIT_C9039
IRI: http://purl.obolibrary.org/obo/NCIT_C94676
IRI: http://purl.obolibrary.org/obo/NCIT_C12892
IRI: http://purl.obolibrary.org/obo/NCIT_C69313
IRI: http://purl.bioontology.org/ontology/ATC/R06AE07
IRI: http://purl.obolibrary.org/obo/NCIT_C26717
IRI: http://purl.obolibrary.org/obo/NCIT_C97132
IRI: http://data.europa.eu/esco/isco/C11
IRI: http://purl.bioontology.org/ontology/ATC/R06AB04
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/38102005
IRI: http://purl.obolibrary.org/obo/NCIT_C122822
IRI: http://purl.obolibrary.org/obo/NCIT_C7405
IRI: http://purl.obolibrary.org/obo/NCIT_C98541
IRI: http://purl.obolibrary.org/obo/NCIT_C3199
IRI: http://purl.bioontology.org/ontology/ATC/N07CA02
IRI: http://purl.bioontology.org/ontology/ATC/N07CA52
IRI: http://purl.bioontology.org/ontology/ATC/N06AB04
IRI: http://purl.bioontology.org/ontology/ATC/L01BB04
IRI: http://purl.bioontology.org/ontology/ATC/L04AA40
IRI: http://data.europa.eu/esco/isco/C91
IRI: http://purl.obolibrary.org/obo/NCIT_C87069
IRI: http://data.europa.eu/esco/isco/C4
IRI: http://purl.bioontology.org/ontology/ATC/N05BA09
IRI: http://purl.bioontology.org/ontology/ATC/N03AE01
IRI: http://purl.bioontology.org/ontology/ATC/B01AC04
IRI: http://purl.obolibrary.org/obo/NCIT_C34486
IRI: http://purl.obolibrary.org/obo/NCIT_C35350
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/447139008
IRI: http://purl.bioontology.org/ontology/ATC/N02AA59
IRI: http://purl.obolibrary.org/obo/NCIT_C116921
IRI: http://purl.obolibrary.org/obo/NCIT_C34695
IRI: http://purl.bioontology.org/ontology/ATC/A11CC05
IRI: http://purl.obolibrary.org/obo/NCIT_C4349
IRI: http://purl.obolibrary.org/obo/NCIT_C4847
IRI: http://purl.obolibrary.org/obo/NCIT_C2955
IRI: http://data.europa.eu/esco/isco/C01
IRI: http://purl.obolibrary.org/obo/NCIT_C34499
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/110469002
IRI: http://purl.obolibrary.org/obo/NCIT_C34360
IRI: http://purl.obolibrary.org/obo/NCIT_C15402
IRI: http://purl.bioontology.org/ontology/ATC/V08
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/232717009
IRI: http://purl.bioontology.org/ontology/ATC/H02
IRI: http://purl.bioontology.org/ontology/ATC/D07
IRI: http://purl.bioontology.org/ontology/ATC/H01AA01
IRI: https://w3id.org/brainteaser/ontology/schema/CortisoneTherapyForRelapse
IRI: http://purl.bioontology.org/ontology/ATC/R05
IRI: http://purl.obolibrary.org/obo/NCIT_C71410
IRI: http://purl.obolibrary.org/obo/NCIT_C171133
IRI: https://w3id.org/brainteaser/ontology/schema/J07BX03
IRI: http://data.europa.eu/esco/isco/C7
IRI: http://purl.obolibrary.org/obo/NCIT_C26769
IRI: http://data.europa.eu/esco/isco/C42
IRI: http://purl.obolibrary.org/obo/NCIT_C3510
IRI: http://purl.bioontology.org/ontology/ATC/B03BA01
IRI: http://purl.bioontology.org/ontology/ATC/L01AA01
IRI: http://purl.obolibrary.org/obo/NCIT_C2978
IRI: http://purl.bioontology.org/ontology/ATC/B01AE07
IRI: http://purl.bioontology.org/ontology/ATC/L04AC01
IRI: http://purl.bioontology.org/ontology/ATC/M03CA01
IRI: http://purl.obolibrary.org/obo/NCIT_C25165
IRI: http://purl.obolibrary.org/obo/NCIT_C122875
IRI: http://ontology.eil.utoronto.ca/GCI/Foundation/GCI-Foundation.owl#decibel
IRI: http://purl.obolibrary.org/obo/NCIT_C49343
IRI: http://purl.bioontology.org/ontology/ATC/H02AB13
IRI: http://purl.obolibrary.org/obo/NCIT_C4786
IRI: http://purl.bioontology.org/ontology/ATC/M05BX04
IRI: http://purl.obolibrary.org/obo/NCIT_C2982
IRI: http://purl.bioontology.org/ontology/ATC/D
IRI: http://purl.bioontology.org/ontology/ATC/G03AC09
IRI: http://purl.bioontology.org/ontology/ATC/G03AA09
IRI: http://purl.bioontology.org/ontology/ATC/N06AX23
IRI: http://purl.bioontology.org/ontology/ATC/H02AB02
IRI: http://purl.obolibrary.org/obo/NCIT_C2985
IRI: http://purl.obolibrary.org/obo/NCIT_C50530
IRI: http://purl.bioontology.org/ontology/ATC/V04
IRI: http://purl.bioontology.org/ontology/ATC/V09
IRI: http://purl.bioontology.org/ontology/ATC/N05BA01
IRI: http://purl.bioontology.org/ontology/ATC/G03AB08
IRI: http://purl.obolibrary.org/obo/NCIT_C1505
IRI: http://purl.bioontology.org/ontology/ATC/A09
IRI: http://purl.bioontology.org/ontology/ATC/L04AX07
IRI: http://purl.bioontology.org/ontology/ATC/J07AF
IRI: http://purl.obolibrary.org/obo/NCIT_C21009
IRI: http://purl.obolibrary.org/obo/NCIT_C2991
IRI: http://purl.bioontology.org/ontology/ATC/C03
IRI: http://purl.obolibrary.org/obo/NCIT_C122419
IRI: http://purl.bioontology.org/ontology/ATC/A03FA03
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/236825001
IRI: http://data.europa.eu/esco/isco/C83
IRI: http://purl.bioontology.org/ontology/ATC/G03AA12
IRI: http://purl.obolibrary.org/obo/OAE_0000890
IRI: http://purl.obolibrary.org/obo/NCIT_C112208
IRI: http://purl.obolibrary.org/obo/OAE_0000891
IRI: http://purl.bioontology.org/ontology/ATC/A02
IRI: http://purl.bioontology.org/ontology/ATC/A05C
IRI: http://purl.bioontology.org/ontology/ATC/A06
IRI: http://purl.bioontology.org/ontology/ATC/A03
IRI: https://w3id.org/brainteaser/ontology/schema/DrugsForMSSymptomaticTherapy
IRI: https://w3id.org/brainteaser/ontology/schema/DrugsForMSTherapySideEffects
IRI: http://purl.bioontology.org/ontology/ATC/R03
IRI: https://w3id.org/brainteaser/ontology/schema/DrugsForOtherDiseases
IRI: http://purl.bioontology.org/ontology/ATC/M05
IRI: http://purl.bioontology.org/ontology/ATC/A10
IRI: http://purl.bioontology.org/ontology/ATC/N06AX21
IRI: http://www.wurvoc.org/vocabularies/om-1.8/duration_of_9_192_631_770_periods_of_the_radiation_corresponding_to_the_transition_between_the_two_hyperfine_levels_of_the_ground_state_of_the_cesium_133_atom
IRI: http://purl.bioontology.org/ontology/ATC/G04CB02
IRI: http://purl.obolibrary.org/obo/NCIT_C80385
IRI: http://purl.bioontology.org/ontology/ATC/P03
IRI: https://w3id.org/brainteaser/ontology/schema/ecunerv
IRI: http://purl.obolibrary.org/obo/NCIT_C3001
IRI: http://data.europa.eu/esco/isco/C74
IRI: http://data.europa.eu/esco/isco/C9
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/55622001
IRI: http://purl.bioontology.org/ontology/ATC/D02
IRI: http://purl.bioontology.org/ontology/ATC/C09AA02
IRI: http://purl.bioontology.org/ontology/ATC/L02
IRI: http://purl.obolibrary.org/obo/NCIT_C3014
IRI: http://purl.obolibrary.org/obo/NCIT_C3134
IRI: http://purl.bioontology.org/ontology/ATC/N06AB10
IRI: http://purl.bioontology.org/ontology/ATC/A02BC05
IRI: http://purl.obolibrary.org/obo/NCIT_C3478
IRI: http://purl.bioontology.org/ontology/ATC/N05BA19
IRI: http://purl.bioontology.org/ontology/ATC/G03AC08
IRI: http://purl.bioontology.org/ontology/ATC/M01AH05
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/300025007
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/118445002
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/428297005
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/700074001
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/370612006
IRI: http://purl.obolibrary.org/obo/NCIT_C79593
IRI: http://purl.obolibrary.org/obo/NCIT_C12401
IRI: http://purl.bioontology.org/ontology/ATC/C10AX09
IRI: http://purl.bioontology.org/ontology/ATC/N07XX07
IRI: http://purl.obolibrary.org/obo/NCIT_C25174
IRI: http://purl.obolibrary.org/obo/NCIT_C34607
IRI: http://purl.obolibrary.org/obo/NCIT_C27020
IRI: http://purl.obolibrary.org/obo/NCIT_C26774
IRI: http://purl.obolibrary.org/obo/NCIT_C26921
IRI: http://purl.obolibrary.org/obo/NCIT_C3041
IRI: http://purl.obolibrary.org/obo/NCIT_C87497
IRI: http://purl.obolibrary.org/obo/NCIT_C3367
IRI: http://purl.bioontology.org/ontology/ATC/L04AA27
IRI: http://purl.obolibrary.org/obo/NCIT_C71411
IRI: http://purl.obolibrary.org/obo/NCIT_C21481
IRI: http://purl.obolibrary.org/obo/NCIT_C78302
IRI: http://purl.bioontology.org/ontology/ATC/N07CA03
IRI: http://purl.bioontology.org/ontology/ATC/N06AB03
IRI: http://purl.bioontology.org/ontology/ATC/N05CD01
IRI: http://purl.bioontology.org/ontology/ATC/R03BA05
IRI: http://data.europa.eu/esco/isco/C94
IRI: http://data.europa.eu/esco/isco/C75
IRI: http://purl.bioontology.org/ontology/ATC/R03AK07
IRI: http://purl.obolibrary.org/obo/NCIT_C84719
IRI: http://purl.obolibrary.org/obo/NCIT_C3245
IRI: http://purl.bioontology.org/ontology/ATC/C03CA01
IRI: http://purl.obolibrary.org/obo/NCIT_C91852
IRI: http://purl.bioontology.org/ontology/ATC/N03AX12
IRI: http://purl.obolibrary.org/obo/NCIT_C26780
IRI: http://purl.obolibrary.org/obo/NCIT_C34632
IRI: http://purl.obolibrary.org/obo/NCIT_C92560
IRI: http://purl.obolibrary.org/obo/NCIT_C26781
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/9429009
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/442338001
IRI: http://data.europa.eu/esco/isco/C41
IRI: http://purl.bioontology.org/ontology/ATC/V06
IRI: http://purl.obolibrary.org/obo/NCIT_C3021
IRI: http://purl.bioontology.org/ontology/ATC/G
IRI: https://orcid.org/0000-0003-4970-4554
IRI: https://orcid.org/0000-0001-9709-6392
IRI: http://purl.bioontology.org/ontology/ATC/L03AX13
IRI: http://purl.obolibrary.org/obo/NCIT_C26782
IRI: http://purl.bioontology.org/ontology/ATC/A10BB09
IRI: http://purl.bioontology.org/ontology/ATC/A10BB12
IRI: http://purl.bioontology.org/ontology/ATC/H02AB
IRI: http://purl.obolibrary.org/obo/NCIT_C71399
IRI: http://purl.obolibrary.org/obo/NCIT_C96573
IRI: http://purl.obolibrary.org/obo/NCIT_C96574
IRI: http://purl.obolibrary.org/obo/NCIT_C71398
IRI: https://orcid.org/0000-0002-5070-2049
IRI: http://purl.bioontology.org/ontology/ATC/G01
IRI: http://data.europa.eu/esco/isco/C73
IRI: http://purl.obolibrary.org/obo/NCIT_C27191
IRI: http://purl.obolibrary.org/obo/NCIT_C12419
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/311801003
IRI: http://purl.obolibrary.org/obo/NCIT_C36282
IRI: http://purl.obolibrary.org/obo/NCIT_C15454
IRI: http://purl.obolibrary.org/obo/NCIT_C34660
IRI: http://purl.obolibrary.org/obo/NCIT_C34661
IRI: http://data.europa.eu/esco/isco/C32
IRI: http://data.europa.eu/esco/isco/C22
IRI: http://purl.obolibrary.org/obo/NCIT_C3079
IRI: http://purl.obolibrary.org/obo/NCIT_C50577
IRI: http://purl.obolibrary.org/obo/NCIT_C50579
IRI: http://purl.obolibrary.org/obo/NCIT_C185645
IRI: http://purl.obolibrary.org/obo/NCIT_C3095
IRI: http://purl.obolibrary.org/obo/NCIT_C3096
IRI: http://purl.bioontology.org/ontology/ATC/J07BC01
IRI: http://purl.obolibrary.org/obo/NCIT_C3098
IRI: http://purl.obolibrary.org/obo/NCIT_C84705
IRI: http://purl.obolibrary.org/obo/NCIT_C34379
IRI: http://purl.obolibrary.org/obo/NCIT_C34685
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/9389005
IRI: http://purl.obolibrary.org/obo/NCIT_C155871
IRI: http://purl.obolibrary.org/obo/NCIT_C71079
IRI: http://purl.obolibrary.org/obo/NCIT_C40197
IRI: http://purl.obolibrary.org/obo/NCIT_C34876
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/414408004
IRI: http://purl.bioontology.org/ontology/ATC/C10AA
IRI: https://www.comunidad.madrid/hospital/gregoriomaranon/
IRI: http://data.europa.eu/esco/isco/C14
IRI: http://purl.bioontology.org/ontology/ATC/C03AA03
IRI: http://purl.bioontology.org/ontology/ATC/D07AA02
IRI: http://purl.bioontology.org/ontology/ATC/H02AB09
IRI: http://purl.bioontology.org/ontology/ATC/P01BA02
IRI: http://purl.obolibrary.org/obo/NCIT_C37967
IRI: http://purl.obolibrary.org/obo/NCIT_C84770
IRI: http://purl.obolibrary.org/obo/NCIT_C3114
IRI: http://purl.obolibrary.org/obo/NCIT_C3117
IRI: http://purl.obolibrary.org/obo/NCIT_C3122
IRI: http://purl.obolibrary.org/obo/NCIT_C3123
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/302870006
IRI: http://purl.obolibrary.org/obo/NCIT_C3124
IRI: http://purl.bioontology.org/ontology/ATC/N05C
IRI: http://purl.obolibrary.org/obo/NCIT_C26800
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/236886002
IRI: http://purl.bioontology.org/ontology/ATC/M01AE01
IRI: http://purl.obolibrary.org/obo/NCIT_C84776
IRI: http://purl.bioontology.org/ontology/ATC/J06
IRI: https://w3id.org/brainteaser/ontology/schema/ImmunoactiveDrugs
IRI: http://purl.bioontology.org/ontology/ATC/J06BA02
IRI: http://purl.bioontology.org/ontology/ATC/L03
IRI: http://purl.bioontology.org/ontology/ATC/L04
IRI: http://purl.bioontology.org/ontology/ATC/M01AB01
IRI: http://purl.obolibrary.org/obo/NCIT_C34726
IRI: http://data.europa.eu/esco/isco/C35
IRI: http://data.europa.eu/esco/isco/C25
IRI: http://www.wurvoc.org/vocabularies/om-1.8/information_capacity_of_one_binary_digit
IRI: http://purl.obolibrary.org/obo/NCIT_C34690
IRI: http://purl.obolibrary.org/obo/NCIT_C3671
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/445968009
IRI: http://purl.obolibrary.org/obo/NCIT_C28286
IRI: https://imm.medicina.ulisboa.pt/
IRI: http://purl.bioontology.org/ontology/ATC/A10AE04
IRI: http://purl.bioontology.org/ontology/ATC/A10AB04
IRI: http://purl.bioontology.org/ontology/ATC/A10A
IRI: http://purl.bioontology.org/ontology/ATC/L03AB02
IRI: http://purl.bioontology.org/ontology/ATC/L03AB07
IRI: http://purl.bioontology.org/ontology/ATC/L03AB08
IRI: http://purl.obolibrary.org/obo/NCIT_C27006
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/73589001
IRI: http://purl.obolibrary.org/obo/NCIT_C34733
IRI: http://purl.obolibrary.org/obo/NCIT_C8366
IRI: http://purl.obolibrary.org/obo/NCIT_C34736
IRI: http://purl.obolibrary.org/obo/NCIT_C84484
IRI: http://purl.obolibrary.org/obo/NCIT_C26805
IRI: http://purl.obolibrary.org/obo/NCIT_C26806
IRI: http://purl.bioontology.org/ontology/ATC/M01AB15
IRI: http://purl.obolibrary.org/obo/NCIT_C9384
IRI: http://purl.obolibrary.org/obo/NCIT_C98967
IRI: http://purl.obolibrary.org/obo/NCIT_C34753
IRI: http://purl.obolibrary.org/obo/NCIT_C142296
IRI: http://data.europa.eu/esco/isco/C93
IRI: http://purl.bioontology.org/ontology/ATC/A06AD11
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/387731002
IRI: http://purl.bioontology.org/ontology/ATC/N03AX09
IRI: http://purl.obolibrary.org/obo/NCIT_C3107
IRI: https://orcid.org/0000-0002-0676-682X
IRI: http://data.europa.eu/esco/isco/C26
IRI: http://data.europa.eu/esco/isco/C34
IRI: http://www.wurvoc.org/vocabularies/om-1.8/length_of_the_path_travelled_by_light_in_vacuum_during_a_time_interval_of_1_299_792_458_of_a_second
IRI: http://purl.bioontology.org/ontology/ATC/C08CA13
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/89743005
IRI: http://purl.obolibrary.org/obo/NCIT_C26816
IRI: http://purl.bioontology.org/ontology/ATC/N03AX14
IRI: http://purl.bioontology.org/ontology/ATC/R06AE09
IRI: http://purl.bioontology.org/ontology/ATC/N04BA02
IRI: http://purl.bioontology.org/ontology/ATC/H03AA01
IRI: http://purl.obolibrary.org/obo/NCIT_C12429
IRI: http://purl.bioontology.org/ontology/ATC/C10
IRI: http://purl.bioontology.org/ontology/ATC/A05B
IRI: http://purl.obolibrary.org/obo/NCIT_C34786
IRI: http://purl.bioontology.org/ontology/ATC/N05BA06
IRI: http://purl.bioontology.org/ontology/ATC/C09CA01
IRI: http://purl.obolibrary.org/obo/NCIT_C12742
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/23344004
IRI: http://purl.obolibrary.org/obo/NCIT_C50643
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/50172003
IRI: http://purl.obolibrary.org/obo/NCIT_C69314
IRI: http://purl.obolibrary.org/obo/NCIT_C48824
IRI: http://purl.obolibrary.org/obo/NCIT_C26823
IRI: http://purl.obolibrary.org/obo/NCIT_C123330
IRI: http://purl.bioontology.org/ontology/ATC/A02AD02
IRI: http://purl.obolibrary.org/obo/NCIT_C9242
IRI: http://purl.obolibrary.org/obo/NCIT_C9305
IRI: http://purl.obolibrary.org/obo/NCIT_C2920
IRI: http://purl.obolibrary.org/obo/NCIT_C7510
IRI: http://data.europa.eu/esco/isco/C1
IRI: http://data.europa.eu/esco/isco/C61
IRI: http://data.europa.eu/esco/isco/C62
IRI: http://www.wurvoc.org/vocabularies/om-1.8/mass_of_the_international_prototype_of_the_kilogram
IRI: http://purl.obolibrary.org/obo/NCIT_C96575
IRI: http://purl.obolibrary.org/obo/NCIT_C96579
IRI: http://purl.bioontology.org/ontology/ATC/D09
IRI: http://purl.obolibrary.org/obo/NCIT_C7570
IRI: http://purl.bioontology.org/ontology/ATC/N05CH01
IRI: http://purl.obolibrary.org/obo/NCIT_C3852
IRI: http://purl.bioontology.org/ontology/ATC/A07EC02
IRI: http://data.europa.eu/esco/isco/C72
IRI: http://purl.bioontology.org/ontology/ATC/A10BA02
IRI: http://purl.bioontology.org/ontology/ATC/L01BA01
IRI: http://purl.bioontology.org/ontology/ATC/L04AX03
IRI: http://purl.bioontology.org/ontology/ATC/H02AB04
IRI: http://purl.obolibrary.org/obo/NCIT_C89715
IRI: http://purl.obolibrary.org/obo/NCIT_C117005
IRI: http://purl.bioontology.org/ontology/ATC/A12
IRI: http://purl.bioontology.org/ontology/ATC/L01DB07
IRI: http://purl.bioontology.org/ontology/ATC/N06AX11
IRI: http://purl.bioontology.org/ontology/ATC/N06BA07
IRI: http://purl.obolibrary.org/obo/NCIT_C112357
IRI: https://www.mondino.it/
IRI: http://purl.obolibrary.org/obo/NCIT_C80310
IRI: http://purl.obolibrary.org/obo/NCIT_C35548
IRI: http://purl.bioontology.org/ontology/ATC/R03DC03
IRI: http://purl.obolibrary.org/obo/NCIT_C92200
IRI: http://purl.obolibrary.org/obo/NCIT_C25189
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/37340000
IRI: http://purl.obolibrary.org/obo/NCIT_C180747
IRI: http://purl.obolibrary.org/obo/NCIT_C3243
IRI: http://purl.obolibrary.org/obo/NCIT_C29888
IRI: http://purl.bioontology.org/ontology/ATC/M03
IRI: http://purl.bioontology.org/ontology/ATC/M
IRI: http://purl.obolibrary.org/obo/NCIT_C60989
IRI: http://purl.obolibrary.org/obo/NCIT_C4345
IRI: http://purl.obolibrary.org/obo/NCIT_C27996
IRI: http://purl.bioontology.org/ontology/ATC/N07BB04
IRI: http://purl.bioontology.org/ontology/ATC/R01
IRI: http://purl.bioontology.org/ontology/ATC/L04AA23
IRI: http://purl.bioontology.org/ontology/ATC/C07AB12
IRI: http://purl.obolibrary.org/obo/NCIT_C13063
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/90460009
IRI: http://purl.obolibrary.org/obo/NCIT_C50667
IRI: http://purl.obolibrary.org/obo/NCIT_C3262
IRI: http://purl.obolibrary.org/obo/NCIT_C71409
IRI: http://purl.bioontology.org/ontology/ATC/N
IRI: http://purl.obolibrary.org/obo/NCIT_C26835
IRI: http://purl.obolibrary.org/obo/NCIT_C6727
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/303011007
IRI: http://purl.bioontology.org/ontology/ATC/C04AE02
IRI: https://orcid.org/0000-0001-9219-6239
IRI: http://purl.obolibrary.org/obo/NCIT_C71408
IRI: http://data.europa.eu/esco/isco/C02
IRI: http://purl.obolibrary.org/obo/NCIT_C3211
IRI: http://purl.obolibrary.org/obo/NCIT_C171457
IRI: http://purl.obolibrary.org/obo/NCIT_C27586
IRI: http://data.europa.eu/esco/isco/C43
IRI: http://purl.obolibrary.org/obo/NCIT_C3283
IRI: http://purl.obolibrary.org/obo/NCIT_C27168
IRI: http://purl.bioontology.org/ontology/ATC/L04AA36
IRI: http://purl.bioontology.org/ontology/ATC/L01FA02
IRI: http://purl.bioontology.org/ontology/ATC/N05AH03
IRI: http://purl.bioontology.org/ontology/ATC/C09CA08
IRI: http://purl.bioontology.org/ontology/ATC/C09DB02
IRI: http://purl.bioontology.org/ontology/ATC/A02BC01
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/21371007
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/428402007
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/428614005
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/107769006
IRI: http://purl.bioontology.org/ontology/ATC/S01
IRI: http://purl.bioontology.org/ontology/ATC/S03
IRI: http://purl.obolibrary.org/obo/NCIT_C157942
IRI: http://purl.obolibrary.org/obo/NCIT_C84950
IRI: http://purl.obolibrary.org/obo/NCIT_C119640
IRI: http://purl.obolibrary.org/obo/NCIT_C3298
IRI: http://purl.bioontology.org/ontology/ATC/A16
IRI: http://data.europa.eu/esco/isco/C44
IRI: http://purl.bioontology.org/ontology/ATC/D11
IRI: http://purl.bioontology.org/ontology/ATC/M09
IRI: http://purl.bioontology.org/ontology/ATC/G02
IRI: http://purl.bioontology.org/ontology/ATC/B06
IRI: http://purl.bioontology.org/ontology/ATC/N07
IRI: http://purl.bioontology.org/ontology/ATC/R07
IRI: http://purl.bioontology.org/ontology/ATC/J07BX
IRI: http://purl.bioontology.org/ontology/ATC/S02
IRI: http://purl.bioontology.org/ontology/ATC/N03AF02
IRI: http://purl.bioontology.org/ontology/ATC/G04BD04
IRI: http://purl.bioontology.org/ontology/ATC/L04AA38
IRI: http://purl.bioontology.org/ontology/ATC/N05AX13
IRI: http://purl.obolibrary.org/obo/NCIT_C3469
IRI: http://purl.bioontology.org/ontology/ATC/H04
IRI: http://purl.obolibrary.org/obo/NCIT_C34889
IRI: http://purl.obolibrary.org/obo/NCIT_C34890
IRI: http://purl.bioontology.org/ontology/ATC/A02BC02
IRI: http://purl.bioontology.org/ontology/ATC/N02BE01
IRI: http://purl.obolibrary.org/obo/NCIT_C26845
IRI: http://purl.bioontology.org/ontology/ATC/N06AB05
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/112902005
IRI: http://purl.obolibrary.org/obo/NCIT_C96581
IRI: http://purl.obolibrary.org/obo/NCIT_C96585
IRI: http://purl.bioontology.org/ontology/ATC/L03AB10
IRI: http://purl.bioontology.org/ontology/ATC/L03AB13
IRI: http://purl.obolibrary.org/obo/NCIT_C12767
IRI: http://purl.obolibrary.org/obo/NCIT_C34911
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/235159007
IRI: http://purl.bioontology.org/ontology/ATC/C09BB04
IRI: http://purl.bioontology.org/ontology/ATC/C04
IRI: http://data.europa.eu/esco/isco/C53
IRI: http://data.europa.eu/esco/isco/C51
IRI: http://purl.obolibrary.org/obo/NCIT_C34922
IRI: http://purl.bioontology.org/ontology/ATC/N03AA02
IRI: http://purl.bioontology.org/ontology/ATC/N03AB02
IRI: http://purl.bioontology.org/ontology/ATC/H01
IRI: http://purl.obolibrary.org/obo/NCIT_C3329
IRI: http://purl.obolibrary.org/obo/NCIT_C26855
IRI: http://data.europa.eu/esco/isco/C8
IRI: http://purl.obolibrary.org/obo/NCIT_C15304
IRI: http://purl.obolibrary.org/obo/NCIT_C3331
IRI: http://purl.obolibrary.org/obo/NCIT_C157959
IRI: http://purl.obolibrary.org/obo/NCIT_C3333
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/882784691000119100
IRI: http://purl.bioontology.org/ontology/ATC/J07BF03
IRI: http://purl.obolibrary.org/obo/NCIT_C117084
IRI: http://purl.bioontology.org/ontology/ATC/N04BC05
IRI: http://purl.bioontology.org/ontology/ATC/B01AC22
IRI: http://purl.bioontology.org/ontology/ATC/N05BA11
IRI: http://purl.bioontology.org/ontology/ATC/H02AB06
IRI: http://purl.bioontology.org/ontology/ATC/H02AB07
IRI: http://purl.bioontology.org/ontology/ATC/N03AX16
IRI: http://purl.bioontology.org/ontology/ATC/D03
IRI: http://purl.obolibrary.org/obo/NCIT_C129933
IRI: http://purl.obolibrary.org/obo/NCIT_C8509
IRI: http://purl.obolibrary.org/obo/NCIT_C85024
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/609637006
IRI: http://data.europa.eu/esco/isco/C13
IRI: http://data.europa.eu/esco/isco/C2
IRI: http://purl.bioontology.org/ontology/ATC/G03AB
IRI: http://purl.bioontology.org/ontology/ATC/A02BX06
IRI: http://purl.obolibrary.org/obo/NCIT_C85026
IRI: http://purl.obolibrary.org/obo/NCIT_C26815
IRI: http://purl.obolibrary.org/obo/NCIT_C85027
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/202708005
IRI: http://purl.bioontology.org/ontology/ATC/C07AA05
IRI: http://data.europa.eu/esco/isco/C54
IRI: http://purl.obolibrary.org/obo/NCIT_C38012
IRI: http://purl.obolibrary.org/obo/NCIT_C3346
IRI: http://purl.obolibrary.org/obo/NCIT_C61277
IRI: http://purl.obolibrary.org/obo/NCIT_C2893
IRI: http://purl.bioontology.org/ontology/ATC/N06
IRI: http://purl.bioontology.org/ontology/ATC/N05
IRI: http://purl.obolibrary.org/obo/NCIT_C78576
IRI: http://purl.obolibrary.org/obo/NCIT_C50713
IRI: http://purl.obolibrary.org/obo/NCIT_C51447
IRI: http://purl.obolibrary.org/obo/NCIT_C34997
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/233935004
IRI: http://purl.bioontology.org/ontology/ATC/N05AH04
IRI: http://purl.obolibrary.org/obo/NCIT_C99039
IRI: http://purl.bioontology.org/ontology/ATC/C09AA05
IRI: http://purl.bioontology.org/ontology/ATC/N06AX18
IRI: http://purl.obolibrary.org/obo/NCIT_C148427
IRI: http://data.europa.eu/esco/isco/C96
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/300009002
IRI: http://purl.obolibrary.org/obo/NCIT_C78593
IRI: http://purl.bioontology.org/ontology/ATC/A10BX02
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/74146001
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/35926005
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/26212005
IRI: http://purl.bioontology.org/ontology/ATC/R
IRI: http://purl.obolibrary.org/obo/NCIT_C26874
IRI: http://purl.obolibrary.org/obo/NCIT_C2884
IRI: http://purl.obolibrary.org/obo/NCIT_C27204
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/20805001
IRI: http://purl.bioontology.org/ontology/ATC/J04AB02
IRI: http://purl.bioontology.org/ontology/ATC/N07XX02
IRI: http://purl.bioontology.org/ontology/ATC/L01XC02
IRI: http://purl.bioontology.org/ontology/ATC/B01AF01
IRI: http://purl.bioontology.org/ontology/ATC/N06DA03
IRI: http://purl.bioontology.org/ontology/ATC/N04BC04
IRI: http://purl.bioontology.org/ontology/ATC/C10AA07
IRI: http://purl.bioontology.org/ontology/ATC/R03CC02
IRI: http://data.europa.eu/esco/isco/C52
IRI: http://purl.obolibrary.org/obo/NCIT_C51605
IRI: http://purl.obolibrary.org/obo/NCIT_C34995
IRI: http://purl.obolibrary.org/obo/NCIT_C94575
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/23056005
IRI: http://data.europa.eu/esco/isco/C31
IRI: http://data.europa.eu/esco/isco/C21
IRI: http://purl.obolibrary.org/obo/NCIT_C19811
IRI: http://purl.bioontology.org/ontology/ATC/L04AC10
IRI: http://purl.obolibrary.org/obo/NCIT_C3020
IRI: http://purl.bioontology.org/ontology/ATC/A10BJ06
IRI: http://purl.bioontology.org/ontology/ATC/S
IRI: http://purl.obolibrary.org/obo/NCIT_C3364
IRI: http://purl.obolibrary.org/obo/NCIT_C70428
IRI: http://purl.bioontology.org/ontology/ATC/N06AB06
IRI: http://purl.obolibrary.org/obo/NCIT_C55098
IRI: http://data.europa.eu/esco/isco/C5
IRI: http://purl.bioontology.org/ontology/ATC/G03
IRI: http://purl.obolibrary.org/obo/NCIT_C35020
IRI: http://purl.obolibrary.org/obo/NCIT_C53458
IRI: http://purl.obolibrary.org/obo/NCIT_C100104
IRI: http://purl.bioontology.org/ontology/ATC/G04BE03
IRI: http://purl.bioontology.org/ontology/ATC/C10AA01
IRI: http://purl.obolibrary.org/obo/NCIT_C35024
IRI: http://purl.bioontology.org/ontology/ATC/L04AA42
IRI: http://purl.obolibrary.org/obo/NCIT_C111202
IRI: http://purl.bioontology.org/ontology/ATC/A10BH01
IRI: http://purl.obolibrary.org/obo/NCIT_C26883
IRI: http://data.europa.eu/esco/isco/C6
IRI: http://purl.obolibrary.org/obo/NCIT_C2921
IRI: http://purl.obolibrary.org/obo/NCIT_C2922
IRI: http://purl.obolibrary.org/obo/NCIT_C4914
IRI: http://purl.obolibrary.org/obo/NCIT_C3884
IRI: http://purl.obolibrary.org/obo/NCIT_C54578
IRI: http://purl.obolibrary.org/obo/NCIT_C25205
IRI: http://purl.obolibrary.org/obo/NCIT_C28381
IRI: http://purl.obolibrary.org/obo/NCIT_C101214
IRI: http://purl.obolibrary.org/obo/NCIT_C85233
IRI: http://purl.obolibrary.org/obo/NCIT_C12464
IRI: http://purl.obolibrary.org/obo/NCIT_C101491
IRI: http://purl.obolibrary.org/obo/NCIT_C35038
IRI: http://data.europa.eu/esco/isco/C81
IRI: https://orcid.org/0000-0003-0362-5893
IRI: http://purl.bioontology.org/ontology/ATC/A01
IRI: http://data.europa.eu/esco/isco/C95
IRI: http://purl.obolibrary.org/obo/NCIT_C3390
IRI: http://purl.obolibrary.org/obo/NCIT_C116585
IRI: http://data.europa.eu/esco/isco/C63
IRI: http://purl.bioontology.org/ontology/ATC/A07EC01
IRI: http://purl.bioontology.org/ontology/ATC/N02CC01
IRI: http://purl.obolibrary.org/obo/NCIT_C12512
IRI: http://purl.bioontology.org/ontology/ATC/V20
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/428490009
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/107784002
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/385487005
IRI: http://purl.obolibrary.org/obo/NCIT_C4876
IRI: http://purl.obolibrary.org/obo/NCIT_C35053
IRI: http://purl.bioontology.org/ontology/ATC/H
IRI: http://purl.bioontology.org/ontology/ATC/G04BE08
IRI: http://purl.bioontology.org/ontology/ATC/S01EE05
IRI: http://purl.bioontology.org/ontology/ATC/L02BA01
IRI: http://purl.bioontology.org/ontology/ATC/G04CA02
IRI: http://purl.obolibrary.org/obo/NCIT_C80020
IRI: http://purl.obolibrary.org/obo/NCIT_C24845
IRI: http://data.europa.eu/esco/isco/C23
IRI: http://data.europa.eu/esco/isco/C3
IRI: http://purl.bioontology.org/ontology/ATC/C09CA07
IRI: http://purl.bioontology.org/ontology/ATC/R03AC03
IRI: http://purl.bioontology.org/ontology/ATC/L04AA31
IRI: http://purl.bioontology.org/ontology/ATC/J06BB02
IRI: http://purl.bioontology.org/ontology/ATC/H01AA02
IRI: http://purl.obolibrary.org/obo/NCIT_C84505
IRI: http://purl.obolibrary.org/obo/NCIT_C35069
IRI: http://purl.bioontology.org/ontology/ATC/V10
IRI: http://www.wurvoc.org/vocabularies/om-1.8/thermodynamic_temperature_of_the_triple_point_of_water
IRI: http://purl.bioontology.org/ontology/ATC/H03BB02
IRI: http://purl.bioontology.org/ontology/ATC/R06AD03
IRI: http://purl.obolibrary.org/obo/NCIT_C100808
IRI: http://purl.obolibrary.org/obo/NCIT_C12894
IRI: http://purl.obolibrary.org/obo/NCIT_C69315
IRI: http://purl.obolibrary.org/obo/NCIT_C171204
IRI: http://purl.obolibrary.org/obo/NCIT_C12799
IRI: http://purl.bioontology.org/ontology/ATC/R02
IRI: http://purl.obolibrary.org/obo/NCIT_C3408
IRI: http://purl.obolibrary.org/obo/NCIT_C28195
IRI: http://purl.obolibrary.org/obo/NCIT_C26893
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/13619001
IRI: http://purl.bioontology.org/ontology/ATC/H03
IRI: http://purl.obolibrary.org/obo/NCIT_C26894
IRI: http://purl.obolibrary.org/obo/NCIT_C99083
IRI: http://purl.bioontology.org/ontology/ATC/G03CX01
IRI: http://purl.bioontology.org/ontology/ATC/B01AC05
IRI: http://purl.obolibrary.org/obo/NCIT_C112833
IRI: http://purl.bioontology.org/ontology/ATC/R03BB04
IRI: http://purl.bioontology.org/ontology/ATC/M03BX02
IRI: http://purl.bioontology.org/ontology/ATC/L04AC07
IRI: http://purl.bioontology.org/ontology/ATC/G04BD07
IRI: http://purl.bioontology.org/ontology/ATC/A13
IRI: http://www.wurvoc.org/vocabularies/om-1.8/tonne
IRI: http://purl.obolibrary.org/obo/NCIT_C35475
IRI: http://purl.bioontology.org/ontology/ATC/M02
IRI: http://purl.bioontology.org/ontology/ATC/N03AX11
IRI: http://purl.obolibrary.org/obo/NCIT_C3418
IRI: http://purl.obolibrary.org/obo/NCIT_C50781
IRI: http://purl.obolibrary.org/obo/NCIT_C15342
IRI: http://purl.bioontology.org/ontology/SNOMEDCT/90199006
IRI: http://purl.bioontology.org/ontology/ATC/L01XC03
IRI: http://purl.bioontology.org/ontology/ATC/N06AX05
IRI: http://purl.bioontology.org/ontology/ATC/N05CD05
IRI: http://purl.obolibrary.org/obo/NCIT_C26952
IRI: http://purl.bioontology.org/ontology/ATC/G04BD09
IRI: http://purl.obolibrary.org/obo/NCIT_C51750
IRI: http://purl.obolibrary.org/obo/NCIT_C100023
IRI: http://purl.obolibrary.org/obo/NCIT_C2986
IRI: http://purl.obolibrary.org/obo/NCIT_C26747
IRI: http://purl.obolibrary.org/obo/NCIT_C96587
IRI: https://www.unipd.it/
IRI: https://www.unito.it/
IRI: http://purl.obolibrary.org/obo/NCIT_C12671
IRI: http://purl.obolibrary.org/obo/NCIT_C115958
IRI: http://purl.obolibrary.org/obo/NCIT_C26905
IRI: http://purl.bioontology.org/ontology/ATC/G04
IRI: http://purl.bioontology.org/ontology/ATC/A05AA02
IRI: http://purl.obolibrary.org/obo/NCIT_C3432
IRI: http://purl.bioontology.org/ontology/ATC/L04AC05
IRI: http://purl.obolibrary.org/obo/NCIT_C3434
IRI: http://purl.bioontology.org/ontology/ATC/J07
IRI: http://purl.bioontology.org/ontology/ATC/J05AB11
IRI: http://purl.bioontology.org/ontology/ATC/N03AG01
IRI: http://purl.bioontology.org/ontology/ATC/C09CA03
IRI: http://purl.bioontology.org/ontology/ATC/V
IRI: http://purl.bioontology.org/ontology/ATC/C05
IRI: http://purl.bioontology.org/ontology/ATC/N06AX16
IRI: http://purl.bioontology.org/ontology/ATC/C08DA01
IRI: http://purl.obolibrary.org/obo/NCIT_C12998
IRI: http://purl.obolibrary.org/obo/NCIT_C38057
IRI: http://purl.bioontology.org/ontology/ATC/N06AX26
IRI: http://purl.bioontology.org/ontology/ATC/N03AG04
IRI: http://purl.obolibrary.org/obo/NCIT_C128405
IRI: http://purl.obolibrary.org/obo/NCIT_C27219
IRI: http://purl.obolibrary.org/obo/NCIT_C114830
IRI: http://purl.bioontology.org/ontology/ATC/A11
IRI: http://purl.obolibrary.org/obo/NCIT_C26915
IRI: http://purl.obolibrary.org/obo/NCIT_C2914
IRI: http://purl.obolibrary.org/obo/NCIT_C35131
IRI: http://purl.bioontology.org/ontology/ATC/B01AA03
IRI: http://purl.bioontology.org/ontology/ATC/N02CC03
IRI: http://purl.bioontology.org/ontology/ATC/N05CF02
IRI: http://purl.bioontology.org/ontology/ATC/N05AF05
[1] Davenport, T., Kalakota, R.: The Potential for Artificial Intelligence in Healthcare. Future Healthc J. 6(2), 94–98 (2019). doi:10.7861/futurehosp.6-2-94
[2] Wang, Y., Wang, L., Rastegar-Mojarad, M., Moon, S., Shen, F., Afzal, N., Liu, S., Zeng, Y., Mehrabi, S., Sohn, S., Liu, H.: Clinical Information Extraction Applications: A Literature Review. J. Biomed. Informatics 77, 34–49 (2018). doi:10.1016/j.jbi.2017.11.011
[3] Cimiano, P., Unger, C., McCrae, J.P.: Ontology-Based Interpretation of Natural Language. Synthesis Lectures on Human Language Technologies. Morgan & Claypool Publishers (2014)
[4] Adams, R.D., Victor, M., Ropper, A.H.: Principles of Neurology. 6th Edition vol. 24. McGraw Hill (1997)
[5] Guazzo, A., Trescato, I., Longato, E., Hazizaj, E., Dosso, D., Faggioli, G., Di Nunzio, G., Silvello, G., Vettoretti, M., Tavazzi, E., et al.: Overview of idpp@ clef 2022: the intelligent disease progression prediction challenge. In: CLEF, pp. 1613–0073 (2022)
[6] Guazzo, A., Trescato, I., Longato, E., Hazizaj, E., Dosso, D., Faggioli, G., Di Nunzio, G.M., Silvello, G., Vettoretti, M., Tavazzi, E., et al.: Intelligent disease progression prediction: overview of idpp@ clef 2022. In: International Conference of the Cross-Language Evaluation Forum for European Languages, pp. 395–422 (2022)
[7] Nunes, S., Sousa, R.T., Serrano, F., Branco, R., Soares, D.F., Martins, A.S., Auletta, E., Castanho, E.N., Madeira, S.C., Aidos, H., et al.: Explaining artificial intelligence predictions of disease progression with semantic similarity. In: CLEF, pp. 1613–0073 (2022)
[8] Gruber, T.R.: A translation approach to portable ontology specifications. Knowledge Acquisition 5(2), 199–220 (1993). doi:10.1006/knac.1993.1008
[9] Berners-Lee, T., Hendler, J., Lassila, O.: The semantic web. Scientific american 284(5), 34–43 (2001)
[10] Noy, N.F., McGuinness, D.L., et al.: Ontology development 101: A guide to creating your first ontology. Stanford knowledge systems laboratory technical report KSL-01-05 (2001)
[11] Feigenbaum, L., Herman, I., Hongsermeier, T., Neumann, E., Stephens, S.: The semantic web in action. Scientific American 297, 64–71 (2008). doi:10.1038/scientificamerican1207-90
[12] Gaspari, M., Saletti, D., Scandellari, C., Stecchi, S.: The aedss application ontology: Enhanced automatic assessment of edss in multiple sclerosis (2005)
[13] Gaspari, M., Saletti, D., Scandellari, C., Stecchi, S.: Refining an automatic EDSS scoring expert system for routine clinical use in multiple sclerosis. IEEE Trans. Inf. Technol. Biomed. 13(4), 501–511 (2009). doi:10.1109/TITB.2008.926498
[14] Esposito, A., De Pietro, G.: An Ontological Approach to Classify Potential Lesion in Patients with Multiple Sclerosis. Technical report (2006)
[15] Alfano, B., Brunetti, A., De Pietro, G., Esposito, A.: An Ontology Approach for Classification of Abnormal White Matter in Patients with Multiple Sclerosis. In: Holzinger, A. (ed.) HCI and Usability for Medicine and Health Care, Third Symposium of the Workgroup Human-Computer Interaction and Usability Engineering of the Austrian Computer Society, USAB 2007, Graz, Austria, November, 22, 2007, Proceedings. Lecture Notes in Computer Science, vol. 4799, pp. 389–402. Springer, ??? (2007). doi:10.1007/978-3-540-76805-0 34. https://doi.org/10.1007/978-3-540-76805-0 34
[16] Esposito, M., De Pietro, G.: An ontology-based fuzzy decision support system for multiple sclerosis. Eng. Appl. Artif. Intell. 24(8), 1340–1354 (2011). doi:10.1016/j.engappai.2011.02.002
[17] Jensen, M., Cox, A.P., Ray, P., Teter, B.E., Weinstock-Guttman, B., Ruttenberg, A., Diehl, A.D.: An Ontological Representation and Analysis of Patient-reported and Clinical Outcomes for Multiple Sclerosis. In: Hogan, W.R., Arabandi, S., Brochhausen, M. (eds.) Proceedings of the 5th International Conference on Biomedical Ontology, ICBO 2014, Houston, Texas, USA, October 8-9, 2014. CEUR Workshop Proceedings, vol. 1327, pp. 52–55. CEUR-WS.org, ??? (2014). http://ceur-ws.org/Vol-1327/icbo2014 paper 44.pdf
[18] Cox, A.P., Jensen, M., Duncan, W., Weinstock-Guttman, B., Szigeti, K., Ruttenberg, A., Smith, B., Diehl, A.D.: Ontologies for the study of neurological disease. In: Towards an Ontology of Mental Functioning (ICBO Workshop), Third International Conference on Biomedical Ontology, (2012)
[19] Jensen, M., Cox, A.P., Chaudhry, N., Ng, M., Sule, D., Duncan, W.D., Ray, P., Weinstock-Guttman, B., Smith, B., Ruttenberg, A., Szigeti, K., Diehl, A.D.: The neurological disease ontology. J. Biomed. Semant. 4, 42 (2013). doi:10.1186/2041-1480-4-42
[20] Jensen, M., Cox, A.P., Smith, B., Diehl, A.D.: Representing disease courses: An application of the neurological disease ontology to multiple sclerosis typology. In: Dumontier, M., Hoehndorf, R., Baker, C.J.O. (eds.) Proceedings of the 4th International Conference on Biomedical Ontology, ICBO 2013, Montreal, Canada, July 7-12, 2013. CEUR Workshop Proceedings, vol. 1060, p. 121. CEUR-WS.org, ??? (2013). http://ceur-ws.org/Vol-1060/icbo2013 submission 69.pdf
[21] Malhotra, A., G ̈undel, M., Rajput, A.M., Mevissen, H.-T., Saiz, A., Pastor, X., Lozano-Rubi, R., Martinez-Lapsicina, E.H., Zubizarreta, I., Mueller, B., et al.: Knowledge retrieval from pubmed abstracts and electronic medical records with the multiple sclerosis ontology. PloS one 10(2), 0116718 (2015)
[22] Pappalardo, F., Russo, G., Pennisi, M., Parasiliti Palumbo, G.A., Sgroi, G., Motta, S., Maimone, D.: The potential of computational modeling to predict disease course and treatment response in patients with relapsing multiple sclerosis. Cells 9(3), 586 (2020)
[23] Alshamrani, R., Althbiti, A., Alshamrani, Y., Alkomah, F., Ma, X.: Model-driven decision making in multiple sclerosis research: Existing works and latest trends. Patterns 1(8), 100121 (2020). doi:10.1016/j.patter.2020.100121
[24] Cardoso, S., Aim ́e, X., Meininger, V., Grabli, D., Mora, L.F.M., Cohen, K.B., Charlet, J.: A modular ontology for modeling service provision in a communication network for coordination of care. In: Ugon, A., Karlsson, D., Klein, G.O., Moen, A. (eds.) Building Continents of Knowledge in Oceans of Data: The Future of Co-Created eHealth - Proceedings of MIE 2018, Medical Informatics Europe, Gothenburg, Sweden, April 24-26, 2018. Studies in Health Technology and Informatics, vol. 247, pp. 890–894. IOS Press (2018). doi:10.3233/978-1-61499-852-5-890. https://doi.org/10.3233/978-1-61499-852-5-890
[25] Cardoso, S., Meneton, P., Aim ́e, X., Meininger, V., Grabli, D., Guezennec, G., Charlet, J.: Use of a modular ontology and a semantic annotation tool to describe the care pathway of patients with amyotrophic lateral sclerosis in a coordination network. PLOS ONE 16(1), 1–19 (2021). doi:10.1371/journal.pone.0244604
[26] Hadzic, M., Chen, M., Dillon, T.S.: Towards the mental health ontology. In: Chen, X., Hu, X., Kim, S. (eds.) 2008 IEEE International Conference on Bioinformatics and Biomedicine, BIBM 2008, Philadephia, Pennsylvania, USA, November 3-5, 2008, pp. 284–288. IEEE Computer Society (2008). doi:10.1109/BIBM.2008.59. https://doi.org/10.1109/BIBM.2008.59
[27] Gibaud, B., Forestier, G., Benoit-Cattin, H., Cervenansky, F., Clarysse, P., Friboulet, D., Gaignard, A., Hugonnard, P., Lartizien, C., Liebgott, H., Montagnat, J., Tabary, J., Glatard, T.: Ontovip: An ontology for the annotation of object models used for medical image simulation. J. Biomed. Informatics 52, 279–292 (2014). doi:10.1016/j.jbi.2014.07.008
[28] Subirats, L., Conesa, J., Armayones, M.: Biomedical holistic ontology for people with rare diseases. International Journal of Environmental Research and Public Health 17(17) (2020). doi:10.3390/ijerph17176038
[29] Golbeck, J., Fragoso, G., Hartel, F.W., Hendler, J.A., Oberthaler, J., Parsia, B.: The national cancer institute’s th ́esaurus and ontology. J. Web Semant. 1(1), 75–80 (2003). doi:10.1016/j.websem.2003.07.007
[30] Sioutos, N., de Coronado, S., Haber, M.W., Hartel, F.W., Shaiu, W.-L., Wright, L.W.: Nci thesaurus: a semantic model integrating cancer-related clinical and molecular information. Journal of Biomedical Informatics 40(1), 30–43 (2007). doi:10.1016/j.jbi.2006.02.013
[31] Donnelly, K.: Snomed-ct: The advanced terminology and coding system for ehealth. Studies in health technology and informatics 121, 279 (2006)
[32] Lee, D., de Keizer, N., Lau, F., Cornet, R.: Literature review of snomed ct use. Journal of the American Medical Informatics Association 21(1), 11–19 (2014)
[33] Chang, E.S., Mostafa, J.: The use of SNOMED ct, 2013-2020: a literature review. J. Am. Medical Informatics Assoc. 28(9), 2017–2026 (2021)
[34] Directorate-General for Employment, Social Affairs and Inclusion: ESCO – European Classification of Skills/Competences, Qualifications and Occupations. Luxembourg: Publications Office of the European Union (2013). doi:10.2767/76494
[35] le Vrang, M., Papantoniou, A., Pauwels, E., Fannes, P., Vandensteen, D., Smedt, J.D.: ESCO: boosting job matching in europe with semantic interoperability. Computer 47(10), 57–64 (2014). doi:10.1109/MC.2014.283
[36] Nahler, G.: anatomical therapeutic chemical classification system (ATC), pp. 8–8. Springer, Vienna (2009). doi:10.1007/978-3-211-89836-9 64. https://doi.org/10.1007/978-3-211-89836-9 64
[37] He, Y., Sarntivijai, S., Lin, Y., Xiang, Z., Guo, A., Zhang, S., Jagannathan, D., Toldo, L., Tao, C., Smith, B.: OAE: The Ontology of Adverse Events. J. Biomed. Semant. 5, 29 (2014). doi:10.1186/2041-1480-5-29
[38] Dahleh, D., Fox, M.S.: An environment ontology for global city indicators (iso 37120). Working paper, Entreprise Integration Laboratory, University of Toronto, 5 King’s College Road, Toronto ON, M5S 3G8 (September 2016). Updated: 30 September 2016
[39] Weibel, S., Kunze, J., Lagoze, C., Wolf, M.: Dublin core metadata for resource discovery (No. rfc2413). Technical report (1998)
[40] Bodenreider, O.: The Unified Medical Language System (UMLS): integrating biomedical terminology. Nucleic Acids Research 32(Database-Issue), 267–270 (2004). doi:10.1093/nar/gkh061
[41] Organization, W.H.: International classification of diseases : [9th] ninth revision, basic tabulation list with alphabetic index. World Health Organization (1978)
[42] Organization, W.H.: ICD-10 : international statistical classification of diseases and related health problems : tenth revision. World Health Organization (2004)
[43] Brown, E.G., Wood, L., Wood, S.: The medical dictionary for regulatory activities (meddra). Drug Safety 20, 109–17 (1999). doi:10.2165/00002018-199920020-00002
[44] Simperl, E.: Reusing ontologies on the semantic web: A feasibility study. Data & Knowledge Engineering [68](10), 905–925 (2009). doi:10.1016/j.datak.2009.02.002
[45] Olsson, T., Barcellos, L.F., Alfredsson, L.: Interactions between genetic, lifestyle and environmental risk factors for multiple sclerosis. Nature Reviews Neurology 13 (2017). doi:10.1038/nrneurol.2016.187
[46] Vasta, R., Chia, R., Traynor, B.J., Chi`o, A.: Unraveling the complex interplay between genes, environment, and climate in als. eBioMedicine 75, 103795 (2022)
[47] Holzinger, A., Biemann, C., Pattichis, C.S., Kell, D.B.: What do we need to build explainable AI systems for the medical domain? CoRR abs/1712.09923 (2017)
[48] Giachelle, F., Irrera, O., Silvello, G.: Medtag: a portable and customizable annotation tool for biomedical documents. BMC Medical Informatics Decis. Mak. 21(1), 352 (2021). doi:10.1186/s12911-021-01706-4
[49] Marchesin, S., Giachelle, F., Marini, N., Atzori, M., Boytcheva, S., Buttafuoco, G., Ciompi, F., Di Nunzio, G.M., Fraggetta, F., Irrera, O., M ̈uller, H., Primov, T., Vatrano, S., Silvello, G.: Empowering digital pathology applications through explainable knowledge extraction tools. Journal of Pathology Informatics 13, 100139 (2022). doi:10.1016/j.jpi.2022.100139
The authors would like to thank Silvio Peroni for developing LODE, a Live OWL Documentation Environment, which is used for representing the Cross Referencing Section of this document and Daniel Garijo for developing Widoco, the program used to create the template used in this documentation.